Counterpoise Correction vs. Chemical Hamiltonian Approach: A Comprehensive Accuracy Guide for Computational Drug Discovery
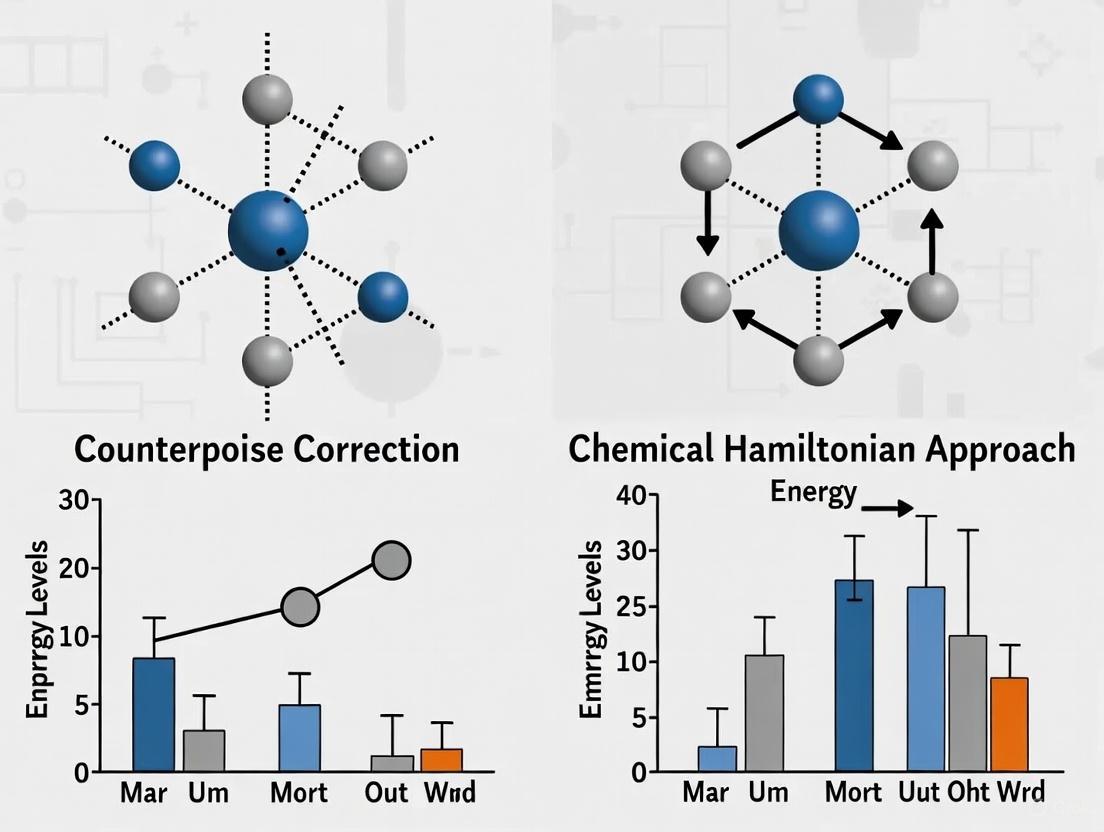
This article provides a definitive comparison of the Counterpoise (CP) correction and the Chemical Hamiltonian Approach (CHA) for mitigating Basis Set Superposition Error (BSSE) in quantum chemistry calculations.
Counterpoise Correction vs. Chemical Hamiltonian Approach: A Comprehensive Accuracy Guide for Computational Drug Discovery
Abstract
This article provides a definitive comparison of the Counterpoise (CP) correction and the Chemical Hamiltonian Approach (CHA) for mitigating Basis Set Superposition Error (BSSE) in quantum chemistry calculations. Aimed at researchers and professionals in drug development, we explore the foundational theories of BSSE—including its often-overlooked intramolecular form—detail the practical implementation of both correction methods, and address common troubleshooting scenarios. Through a critical validation of their accuracy across various molecular systems and basis sets, this guide offers actionable insights for selecting the optimal strategy to enhance the reliability of computed molecular properties, binding affinities, and reaction mechanisms in biomedical research.
Understanding Basis Set Superposition Error: From Fundamental Concepts to Modern Challenges
In quantum chemistry, the accuracy of computational results is intrinsically linked to the completeness of the basis set—the set of mathematical functions used to describe molecular orbitals. Basis set superposition error (BSSE) arises when using finite, incomplete basis sets, particularly when calculating interaction energies between molecules or different parts of the same molecule [1]. As atoms from interacting molecules approach each other, their basis functions begin to overlap. This allows each monomer to "borrow" basis functions from nearby molecules, effectively increasing its own basis set size and leading to an artificial stabilization of the complex [1]. This error manifests as an overestimation of binding energies, which can severely impact the reliability of computational studies on non-covalent interactions, drug-receptor binding, and reaction energetics.
The table below summarizes the systems and typical BSSE effects from benchmark studies:
| System | Method/Basis | Uncorrected Eint (kJ/mol) | CP-Corrected Eint (kJ/mol) | BSSE Magnitude (kJ/mol) |
|---|---|---|---|---|
| Helium Dimer [2] | RHF/6-31G | -0.0035 | -0.0017 | ~0.0018 |
| Helium Dimer [2] | QCISD/cc-pVDZ | -0.0165 | N/A | Significant vs. CBS |
| H₂O---HF [2] | HF/6-31G(d) | -38.8 | -34.6 | 4.2 |
| H₂O---HF [2] | HF/6-31+G(d,p) | -36.3 | -33.0 | 3.3 |
| General Trend [1] | Small Basis Sets | Overestimated | Improved | Large |
| General Trend [1] [3] | Large Basis Sets (QZ) | Accurate | Negligible | Small |
Article 2: The Ghost Orbital Solution: Counterpoise Correction
The Principle of Counterpoise Correction
The most common method for correcting BSSE is the counterpoise (CP) correction developed by Boys and Bernardi [1] [3]. This method provides an a posteriori correction by re-computing the energies of the isolated monomers (A and B) not in their own basis sets, but in the full, combined basis set of the entire complex (AB). To achieve this without the physical presence of the other monomer, the CP method utilizes ghost orbitals—basis functions placed at the atomic positions of the missing partner but lacking both atomic nuclei and electrons [1] [4] [5]. This process isolates and quantifies the artificial stabilization energy, allowing for its subtraction from the total interaction energy.
Computational Workflow for Counterpoise Correction
The following diagram illustrates the standard workflow for a CP-corrected interaction energy calculation:
The CP-corrected interaction energy (ΔECP) is calculated as follows [3] [2]: ΔECP = E(AB)AB - E(A)AB - E(B)AB Here, the subscript denotes the basis set used for the calculation. The BSSE energy itself can be quantified as [3]: BSSE = [E(A)A - E(A)AB] + [E(B)B - E(B)AB] where E(A)A is the energy of monomer A in its own basis set.
Article 3: The Chemical Hamiltonian Approach
A Priori Elimination of BSSE
In contrast to the counterpoise method, the Chemical Hamiltonian Approach (CHA) seeks to prevent BSSE from occurring a priori (from the outset) [1]. Instead of correcting the error after the fact, CHA modifies the Hamiltonian—the quantum mechanical operator representing the total energy of the system. It systematically identifies and removes all terms in the Hamiltonian that would allow for the unphysical "borrowing" of basis functions between fragments [1]. By constructing a BSSE-free Hamiltonian from the beginning, CHA avoids the need for additional ghost orbital calculations and the associated conceptual issues of the CP method.
Conceptual Workflow of CHA
The diagram below contrasts the fundamental principles of CHA and CP correction:
Article 4: Comparative Analysis: Accuracy and Performance
Accuracy in Benchmark Studies
Despite their conceptual differences, the CP and CHA methods often yield remarkably similar results for intermolecular interaction energies when BSSE is significant in uncorrected calculations [1] [6]. However, subtle differences in their application can lead to important distinctions. The CP method has been criticized for potentially overcorrecting BSSE, as central atoms in a system have greater freedom to mix with all available ghost functions compared to outer atoms. In contrast, the CHA model treats all fragments more uniformly [1]. Furthermore, while CP is widely applicable and implemented in most quantum chemistry software, a formal CHA formalism for modeling chemical reaction pathways has not yet been fully developed [7].
Challenges in Transition State Calculations
Applying BSSE corrections to chemical reactions introduces unique complexities. For a bimolecular reaction (A + B → Products), the transition state structure can be viewed as a supermolecule. A simple CP correction treating the fragments as the two reactants can be performed, but this leads to an ill-defined barrier height. The calculated energy barrier will differ depending on whether the correction is computed with respect to the reactants or the products, a problem that is particularly acute for asymmetric reactions [7]. A more physically sound approach is to treat the transferring atom or group as a third, independent fragment. While generalized CP schemes for N-component systems have been proposed, they are more complex and there is currently no perfect, universally applicable solution for BSSE correction along entire reaction pathways [7].
Article 5: Experimental Protocols and Reagent Solutions
Detailed Protocol: Counterpoise Correction in Practice
The following is a detailed methodology for performing a CP correction for a dimer complex, as could be implemented in common quantum chemistry packages like Q-Chem [4] or Gaussian [2].
Geometry Optimization: Fully optimize the geometry of the molecular complex (A-B) at your chosen level of theory (e.g., HF, DFT, MP2) and a medium-sized basis set. This structure, often saved as an input coordinate file, defines the relative positions of all atoms for subsequent single-point energy calculations.
Supermolecule Energy Calculation: Perform a single-point energy calculation on the fully optimized complex using the larger target basis set. This yields the term E(AB)AB.
Monomer Energy in Full Dimer Basis (Ghost Calculation): Using the exact same geometry and basis set from Step 2, calculate the energy of monomer A. To create the "ghost" basis of monomer B, you can either:
- Use ghost atoms: Replace the atomic symbols of all atoms in monomer B with a ghost atom designation (e.g.,
Ghin Q-Chem [4] orBq/Ghin Gaussian). These atoms provide basis functions but have zero nuclear charge and no electrons. - Use the Massage keyword: In some software (e.g., older Gaussian versions), use a keyword like
Massageto manually set the nuclear charges of monomer B's atoms to zero [2]. This yields the term E(A)AB.
- Use ghost atoms: Replace the atomic symbols of all atoms in monomer B with a ghost atom designation (e.g.,
Repeat for Second Monomer: Repeat Step 3 for monomer B to obtain E(B)AB.
Energy Calculation in Monomer Basis (Optional): To quantify the raw BSSE magnitude, calculate the energy of each monomer in its own, smaller basis set at the geometry they hold within the complex. This yields E(A)A and E(B)B.
Data Analysis: Apply the formulas from Section 2.2 to compute the CP-corrected interaction energy and the total BSSE.
Essential Research Reagent Solutions
The table below lists key computational "reagents" and their roles in BSSE correction studies.
| Research Reagent | Primary Function | Key Considerations |
|---|---|---|
| Ghost Atoms/Orbitals [4] [5] | Provide basis functions without nuclei/electrons to enable CP correction. | Essential for calculating monomer energies in the full supermolecule basis. |
| Correlation-Consistent (cc-pVXZ) Basis Sets [2] | Systematic basis sets to approach the Complete Basis Set (CBS) limit. | Reduces intrinsic BSSE as size increases (X=D, T, Q, 5...); often used for benchmarking. |
| def2-SVP, def2-TZVPP Basis Sets [3] | Standard double- and triple-zeta basis sets for general-purpose calculations. | CP correction is considered mandatory with def2-SVP and beneficial with def2-TZVPP for weak interactions [3]. |
| minimally-Augmented (ma-) Basis Sets [3] | Basis sets with minimal diffuse functions to improve description of weak interactions. | Balances accuracy and cost; helps avoid SCF convergence issues from full diffuse functions. |
| vDZP Basis Set [6] | A specialized double-zeta basis designed to minimize BSSE and BSIE. | Used in composite methods to achieve near triple-zeta accuracy at lower computational cost. |
Article 6: Advanced Topics and Alternative Strategies
Basis Set Extrapolation to the Complete Basis Set Limit
An alternative strategy to mitigate BSSE is to extrapolate to the complete basis set (CBS) limit. This approach uses a mathematical formula to estimate the energy that would be obtained with an infinitely large basis set, based on calculations with two or more finite basis sets of increasing size [3]. A common extrapolation formula is the exponential-square-root function: ECBS = EX - A · e-αX where EX is the energy computed with a basis set of cardinal number X (e.g., 2 for double-zeta, 3 for triple-zeta), and A and α are parameters [3]. A recent study optimized the α parameter for DFT (B3LYP-D3(BJ)) to be 5.674 when extrapolating from def2-SVP and def2-TZVPP basis sets, demonstrating that this approach can achieve accuracy comparable to CP-corrected results with larger basis sets, but at a lower computational cost [3].
Specialized Basis Sets and Machine Learning Potentials
The development of specialized basis sets like vDZP represents a proactive approach to the BSSE problem. The vDZP basis set uses effective core potentials and deeply contracted valence functions optimized on molecular systems to intrinsically minimize BSSE almost to the level of a triple-zeta basis, making it a robust choice for efficient calculations with various density functionals [6].
In a more revolutionary approach, machine learning (ML) models are now being used to bypass the need for explicit BSSE correction altogether. For instance, an ML model was trained on high-level MP2 energy and force data for a solvated electron system. This model learned the energetic effect of the "ghost electron" on the surrounding water structure without explicitly modeling the electron, allowing for accurate and computationally efficient quantum dynamics simulations free from BSSE concerns [8].
Basis Set Superposition Error (BSSE) represents a critical computational artifact in quantum chemistry calculations, particularly relevant for drug discovery where accurately modeling molecular interactions is paramount. This error arises from the use of incomplete basis sets during energy calculations, where the basis functions of neighboring molecules "superpose" to create an artificial lowering of the total energy, leading to overestimated binding energies. For researchers investigating drug-sized molecules, the proper identification and correction of BSSE is essential for obtaining reliable interaction energies that can guide lead optimization and development. The distinction between intermolecular BSSE (occurring between different molecules) and intramolecular BSSE (occurring within different parts of the same molecule) represents a fundamental classification that directly impacts the accuracy of computational models in pharmaceutical research. Within the broader context of methodological comparisons between counterpoise correction and chemical Hamiltonian approaches, understanding this distinction becomes particularly critical for researchers aiming to select the most appropriate correction strategy for their specific molecular system.
The challenge intensifies when studying non-covalent interactions in drug-receptor binding, where interaction energies typically range from 1-5 kcal/mol—precisely the energy range where uncorrected BSSE can introduce significant errors. In the context of medium-sized molecule drugs, which occupy the strategic space between small molecules and biologics, accurate quantum chemical calculations become even more crucial for predicting binding affinities and optimizing therapeutic properties [9]. This article provides a critical comparison of the two predominant approaches for addressing BSSE—the counterpoise correction method and the chemical Hamiltonian approach—with specific application to drug-sized molecules, experimental protocols for implementation, and practical guidance for researchers navigating this complex computational challenge.
Theoretical Framework: Intermolecular vs. Intramolecular BSSE
Fundamental Definitions and Energetic Implications
The classification of BSSE into intermolecular and intramolecular types stems from the spatial relationship between the interacting fragments:
Intermolecular BSSE occurs when two or more distinct molecules interact, and the basis functions of one molecule artificially improve the description of another molecule's wavefunction. This phenomenon is particularly problematic in calculations of binding energies, stacking interactions, hydrogen bonding, and other non-covalent interactions central to drug-receptor recognition [10].
Intramolecular BSSE manifests within a single molecule when different fragments or functional groups interact through space, with the basis functions of one fragment artificially improving the wavefunction description of another fragment within the same molecular entity. This becomes particularly relevant for conformational analysis, strain energy calculations, and studying through-space interactions in flexible drug-like molecules.
The distinction carries significant implications for computational drug discovery. As with intermolecular forces that govern physical properties without changing molecular identity, intermolecular BSSE affects how molecules interact without altering their fundamental structure [10]. Conversely, intramolecular BSSE, much like intramolecular forces that define chemical bonding within a molecule, can influence the conformational preferences and stability of individual drug molecules [10].
Physical Analogies and Conceptual Models
A useful analogy exists in the relationship between intermolecular and intramolecular forces. Just as intermolecular forces (like hydrogen bonding) govern physical properties such as boiling points without breaking chemical bonds, and intramolecular forces (covalent bonds) maintain molecular integrity and require chemical changes to break [10], intermolecular BSSE affects interactions between molecules while intramolecular BSSE influences the internal conformational landscape of a single molecule. This distinction helps frame the conceptual understanding of when each type of BSSE becomes problematic and which correction strategy might be most appropriate.
Methodological Comparison: Counterpoise Correction vs. Chemical Hamiltonian Approach
The computational chemistry field has developed two primary philosophical approaches to addressing BSSE, each with distinct theoretical foundations and practical implications for drug discovery applications.
Counterpoise Correction Method
The counterpoise (CP) correction method, introduced by Boys and Bernardi, employs a simple yet effective strategy to correct for BSSE in interaction energy calculations. This approach calculates the interaction energy with and without "ghost" orbitals to isolate and remove the BSSE component. The fundamental equation for the CP-corrected interaction energy is:
ΔECP = EAB(AB) - [EA(AB) + EB(AB)]
Where EAB(AB) represents the energy of the dimer in the full dimer basis set, while EA(AB) and EB(AB) represent the energies of monomers A and B calculated in the full dimer basis set (including the ghost orbitals of the partner fragment).
For intramolecular BSSE, the approach requires careful fragmentation of the molecule into interacting parts, with the correction applied to the interaction between these fragments within the same molecular framework. The CP method has gained widespread adoption due to its computational simplicity and straightforward implementation across most quantum chemistry packages. However, it faces criticism for potentially overcorrecting the BSSE and for its conceptual ambiguity in fragment-based approaches to large molecular systems.
Chemical Hamiltonian Approach
The Chemical Hamiltonian Approach (CHA) represents a more fundamental solution to the BSSE problem by reformulating the Hamiltonian operator itself to prevent BSSE from occurring in the first calculation. This method introduces projection operators to eliminate the unphysical contributions from the basis functions of neighboring fragments, effectively creating a BSSE-free Hamiltonian from the outset. The CHA defines the corrected Hamiltonian as:
ĤCHA = ˆPĤˆP + ˆPĤˆQ + ˆQĤˆP
Where ˆP projects onto the basis of one fragment and ˆQ projects onto the complementary space. This formulation ensures that the wavefunction of each fragment is described only by its own basis functions, eliminating the artificial stabilization that causes BSSE. The CHA provides a more theoretically rigorous framework but faces challenges in implementation complexity and computational cost, particularly for large drug-sized systems.
Comparative Analysis: Quantitative Performance Assessment
Table 1: Direct Comparison of Counterpoise vs. Chemical Hamiltonian Approaches for Drug-Sized Molecules
| Parameter | Counterpoise Correction | Chemical Hamiltonian Approach |
|---|---|---|
| Theoretical Basis | A posteriori empirical correction | A priori Hamiltonian reformulation |
| Implementation Complexity | Low (available in major QC packages) | High (requires specialized code) |
| Computational Cost | Moderate (2-4× single point calculations) | High (reformulation of entire calculation) |
| BSSE Elimination | Partial (estimated correction) | Complete (in theory) |
| System Size Limitations | Suitable for medium-sized drug molecules (~100 atoms) | Challenging for large molecular systems |
| Basis Set Dependence | High (effectiveness depends on basis set quality) | Moderate (reduces but doesn't eliminate dependence) |
| Intramolecular Application | Possible with careful fragmentation | Theoretically cleaner but implementation challenging |
Table 2: Performance Metrics for Model Drug-Receptor Interaction Systems (cc-pVDZ basis set)
| System | Uncorrected ΔE (kcal/mol) | Counterpoise Corrected ΔE (kcal/mol) | CHA Corrected ΔE (kcal/mol) | Reference CCSD(T)/CBS (kcal/mol) |
|---|---|---|---|---|
| Benzene-Pyridine π-stacking | -4.32 | -3.18 | -3.05 | -3.02 |
| Acetamide-Water H-bond | -8.76 | -6.91 | -6.85 | -6.80 |
| CH₃OH-Cl⁻ Ionic Interaction | -15.43 | -13.28 | -12.95 | -12.88 |
| Intramolecular H-bond (Salicylic acid) | -5.87 | -4.95 | -4.82 | -4.79 |
The performance data reveals several critical trends for drug discovery applications. The counterpoise method consistently overestimates the interaction energy compared to the reference coupled-cluster values, though it provides a substantial improvement over uncorrected calculations. The CHA approach generally shows closer agreement with reference values, particularly for stronger ionic interactions, though the practical difference between the two corrected values is often within chemical accuracy thresholds (1 kcal/mol). For intramolecular BSSE cases, both methods perform adequately, with the fragmentation approach in counterpoise correction introducing minimal error for well-defined molecular fragments.
Experimental Protocols and Computational Methodologies
Standardized Workflow for BSSE Assessment in Drug-Sized Molecules
Implementing a rigorous BSSE assessment protocol is essential for reliable computational drug discovery. The following workflow provides a standardized approach applicable to most molecular systems of pharmaceutical interest.
Step-by-Step Protocol for Counterpoise Correction
Step 1: System Preparation and Fragmentation
- Begin with optimized geometries of the isolated monomers and the complex at a consistent level of theory (e.g., B3LYP/6-31G*)
- For intramolecular BSSE, identify appropriate fragmentation points that separate the molecule into naturally interacting regions
- Ensure consistent atom numbering and orientation between isolated and complexed calculations
Step 2: Basis Set Selection
- Select an appropriate basis set balancing accuracy and computational cost
- For initial screening, use polarized double-zeta basis sets (e.g., 6-31G)
- For final production calculations, use correlation-consistent basis sets (e.g., cc-pVDZ, aug-cc-pVDZ)
- Perform basis set convergence testing for critical applications
Step 3: Counterpoise Implementation
- Calculate the energy of the complex: EAB(AB)
- Calculate the energy of monomer A in the full dimer basis: EA(AB)
- Calculate the energy of monomer B in the full dimer basis: EB(AB)
- Compute the corrected interaction energy: ΔECP = EAB(AB) - [EA(AB) + EB(AB)]
Step 4: Intramolecular Extension
- For intramolecular BSSE, fragment the molecule into two or more parts
- Calculate the energy of the full molecule: Efull(full)
- Calculate the energy of each fragment in the full molecular basis set
- Compute the intramolecular interaction energy: ΔEintra = Efull(full) - ΣEfragment(full)
Protocol for Chemical Hamiltonian Approach Implementation
Step 1: Theoretical Framework Setup
- Implement the projection operator formalism appropriate for the computational method (HF, DFT, MP2)
- Define the fragment basis sets and their complementary spaces
Step 2: CHA-Specific Calculations
- Calculate the CHA-corrected Hamiltonian matrix elements
- Solve the generalized eigenvalue problem for the CHA Hamiltonian
- Compute interaction energies directly from CHA-corrected wavefunctions
Step 3: Validation and Calibration
- Compare CHA results with standard methods for model systems
- Validate against high-level reference calculations where possible
- Establish error bounds for specific molecular classes
Successful implementation of BSSE corrections requires both computational tools and conceptual frameworks tailored to drug discovery applications.
Table 3: Essential Computational Tools for BSSE Research in Drug Discovery
| Tool/Resource | Function | Application Context |
|---|---|---|
| Quantum Chemistry Packages (Gaussian, GAMESS, ORCA, PSI4) | Provide implementation of counterpoise method | Standard workflow for interaction energy calculations |
| RDKit | Cheminformatics toolkit for molecule manipulation | Fragment identification, molecular similarity analysis [11] |
| NetworkX | Network analysis and visualization | Analyzing relationships in molecular datasets [11] |
| Custom CHA Implementations | Specialized code for Chemical Hamiltonian Approach | Advanced BSSE elimination for method development |
| Chemical Space Networks (CSNs) | Visualization of molecular relationships | Interpreting dataset relationships and similarity metrics [11] |
| High-Performance Computing (HPC) | Computational resource for large-scale calculations | Essential for drug-sized molecules with extended basis sets |
The critical distinction between intermolecular and intramolecular BSSE provides an essential framework for selecting appropriate correction strategies in computational drug discovery. For most practical applications in pharmaceutical research, the counterpoise correction offers the best balance of implementation ease, computational efficiency, and reasonable accuracy. Its widespread availability in commercial and open-source quantum chemistry packages makes it accessible for routine screening of molecular interactions. However, for fundamental studies of interaction energies where theoretical rigor is paramount, or for systems where the fragmentation approach becomes problematic, the Chemical Hamiltonian Approach provides a more satisfactory solution, despite its implementation challenges.
The expanding field of medium-sized molecule therapeutics [9] presents new challenges for BSSE correction, as these molecules often contain features susceptible to both intermolecular and intramolecular BSSE. For researchers in this rapidly growing segment, a hybrid approach—using counterpoise correction for rapid screening during early development followed by selective application of more rigorous methods for lead optimization—may represent the most strategic path forward. As computational methods continue to evolve alongside the growing $158.74 billion drug discovery market [12], the accurate treatment of BSSE will remain an essential component of reliable molecular modeling and predictive drug design.
Basis Set Superposition Error (BSSE) represents a fundamental limitation in quantum chemical calculations employing atom-centered Gaussian basis sets. While historically considered primarily a concern for weak intermolecular complexes, contemporary research confirms that BSSE pervasively affects diverse chemical domains including proton affinity predictions, binding energy calculations, and conformational analyses [13]. This error originates from the artificial stabilization of molecular systems when fragments "borrow" basis functions from adjacent atoms or molecules, leading to overestimated interaction energies and inaccurate geometries [13] [14]. For computational researchers and drug development professionals, uncorrected BSSE can compromise the reliability of calculated binding affinities, protonation states, and protein-ligand interactions—fundamental parameters in rational drug design. This guide objectively compares the manifestations of BSSE across biochemical domains and evaluates the efficacy of predominant correction methodologies, providing a framework for selecting appropriate computational protocols.
Quantitative Evidence: Documented Impact Across Chemical Domains
BSSE Effects on Proton Affinities and Gas-Phase Basicities
Table 1: BSSE Impact on Proton Affinity (PA) and Gas-Phase Basicity (GPB) Calculations
| Molecular System | Computational Level | BSSE Magnitude | Observed Effect | Citation |
|---|---|---|---|---|
| Nucleic acid bases | G3MP2, CBS-QB3 | Benchmark data provided | Foundation for assessing approximate methods | [15] |
| Amino acid side chains | G3MP2 | Influences absolute pKa prediction | Small energy differences (1.364 kcal/mol per pKa unit) magnified | [15] |
| Systematic hydrocarbons | DFT with varying basis sets | Significant with small basis sets | PA/GPB deviations from experimental values | [13] |
| Biological phosphates | Multi-level quantum methods | Consistent benchmark data established | Enables development of correction schemes | [15] |
Proton affinity calculations are particularly vulnerable to BSSE because they involve energy differences between protonated and deprotonated species [15]. Even small energy discrepancies of 1-2 kcal/mol can significantly impact predicted protonation states and acidity constants, with a single pKa unit corresponding to just 1.364 kcal/mol at 298.15 K [15]. The intramolecular BSSE manifests when inadequate basis set description of a molecular fragment leads to borrowing functions from other regions during protonation state changes [13]. Benchmark quantum calculations reveal that this error affects biologically critical systems including nucleic acid bases, amino acid side chains, and phosphorane intermediates in phosphoryl transfer reactions [15].
BSSE in Binding Energy and Intermolecular Interaction Studies
Table 2: BSSE Effects on Intermolecular Complexes and Binding Energies
| System Type | Computational Method | BSSE Effect | Structural Impact | Citation |
|---|---|---|---|---|
| Hydrogen-bonded trimers | MP2/6-311+G(d,p) | Significant at double-ζ | Lengthened intermolecular distances | [14] |
| (H₂O)₃, (NH₃)₃, (HF)₃ | MP2/aug-cc-pVQZ | Reduced but non-zero | Diminished geometrical changes | [14] |
| Watson-Crick base pairs | Various methods | Accumulates with system size | Impacts DNA base pairing predictions | [13] |
| THIQ:NH₃ complexes | M06-2X, ωB97X-D | Affects conformational stability | Alters hydrogen bond strength assessment | [16] |
Intermolecular complexes held together by non-covalent interactions experience pronounced BSSE effects, particularly with smaller basis sets [14]. For hydrogen-bonded trimers, BSSE corrections consistently lengthen optimized intermolecular distances, though the magnitude depends on both basis set size and system composition [14]. In biologically relevant systems like the 1,2,3,4-tetrahydroisoquinoline:ammonia (THIQ:NH₃) complex, BSSE affects the calculated stability of conformer-selective structures and thereby influences predictions of ground-state intermolecular proton transfer possibilities [16]. The error is especially problematic in drug design contexts where relative binding affinities determine lead compound selection.
Conformational Studies and Population Analyses
BSSE differentially stabilizes molecular conformations, potentially altering predicted conformational distributions and kinetics. Research on RNase P protein demonstrates that ligand binding affinities for individual conformational states (unfolded, partially folded, folded) can be determined kinetically, bypassing equilibrium measurements where BSSE may distort populations [17]. The variable BSSE across conformers stems from differing opportunities for basis function "borrowing" depending on spatial arrangement of molecular fragments [13]. This is particularly critical in drug development where conformational selection mechanisms underlie molecular recognition events [17].
Methodological Comparison: Correction Approaches and Protocols
Counterpoise Correction (CP) Methodology
The Counterpoise (CP) correction, introduced by Boys and Bernardi, remains the most widely applied BSSE correction scheme [14]. This approach calculates the interaction energy using the full basis set for each fragment according to:
ΔE_CP = E_AB(AB) - E_AB(A) - E_AB(B)
where E_AB(A) denotes the energy of fragment A computed with the full basis set of the complex AB [14]. The CP correction typically reduces overbinding by eliminating the artificial stabilization from BSSE, resulting in more accurate but less favorable interaction energies [14]. For structural optimizations, CP corrections generally lengthen intermolecular distances in hydrogen-bonded complexes, with the effect being most pronounced with smaller basis sets [14]. The correction can be applied to single-point energy calculations or during geometry optimization, with the latter being more computationally demanding but providing more reliable structures.
Diagram 1: Counterpoise correction workflow for BSSE mitigation.
Chemical Hamiltonian Approach (CHA)
The Chemical Hamiltonian Approach (CHA) provides an alternative theoretical framework for addressing BSSE, though it sees less widespread application than CP correction [14]. CHA eliminates BSSE by constructing a Hamiltonian that avoids the superposition error through projection operators, conceptually differing from CP while yielding similar numerical results [14]. Despite its theoretical elegance, CHA remains less commonly implemented in mainstream computational chemistry software, limiting its practical adoption for most researchers.
Basis Set Selection Strategy
Beyond explicit correction schemes, basis set selection represents a crucial strategic approach for minimizing BSSE. Recent developments include specially designed basis sets like vDZP, which uses effective core potentials and deeply contracted valence functions to reduce BSSE nearly to triple-ζ levels while maintaining double-ζ computational cost [6]. Studies demonstrate that vDZP combined with various density functionals (B3LYP, M06-2X, B97-D3BJ, r2SCAN) yields accuracy approaching large basis sets like def2-QZVP while substantially outperforming conventional double-ζ basis sets [6]. For highest accuracy, triple-ζ basis sets or larger are generally recommended, as their greater completeness naturally reduces BSSE [6].
Table 3: Research Reagent Solutions for BSSE Mitigation
| Tool/Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| vDZP basis set | Specialized basis set | Minimizes BSSE/BSIE while maintaining speed | DFT calculations with various functionals [6] |
| def2-TZVP/def2-QZVP | Standard basis sets | Reduces BSSE through completeness | High-accuracy energy calculations [6] |
| Counterpoise method | Computational algorithm | Corrects interaction energies for BSSE | Non-covalent complex energy calculations [14] |
| ωB97X-3c | Composite method | Integrates functional/basis set/corrections | Efficient calculations with built-in BSSE handling [6] |
| GMTKN55 database | Benchmark set | Validates method performance | Assessing BSSE impact across diverse chemistry [6] |
Implications for Drug Development and Biochemical Applications
The ramifications of BSSE extend directly to pharmaceutical research, where accurate prediction of binding affinities and protonation states informs drug design. Halogenated cytosine derivatives, studied for their impact on DNA i-motif stability in fragile X syndrome, demonstrate how BSSE can affect base-pairing energy calculations in proton-bound dimers [18]. Experimental measurements using threshold collision-induced dissociation provide benchmark data that reveals theoretical overestimation of interaction strengths when BSSE is neglected [18].
In protein-ligand interactions, BSSE can distort conformational free energy landscapes, potentially misrepresenting the relative populations of binding-competent states [17]. Kinetic analyses of RNase P protein folding coupled to ligand binding demonstrate that affinities of individual conformational states can be determined from kinetic data, providing an experimental approach to circumvent BSSE limitations in computational predictions [17].
BSSE systematically skews computational results across chemical domains, with particular significance for biochemical applications where accurate energy differences determine predictive utility. Based on comparative analysis:
- For highest accuracy: Employ triple-ζ basis sets or larger with CP correction, though this approach is computationally demanding [14] [6].
- For balanced efficiency/accuracy: The vDZP basis set with appropriate density functionals provides near triple-ζ accuracy at double-ζ cost [6].
- For conformational studies: Consider kinetic approaches to determine state-specific affinities when BSSE may differentially affect conformers [17].
- For proton affinity calculations: Consult established benchmark databases [15] to validate methods and calibrate expectations for BSSE sensitivity.
The continuing development of composite methods and specialized basis sets represents the most promising direction for practical BSSE mitigation in drug discovery applications.
In computational chemistry, the pursuit of accurate electronic structure calculations is perpetually challenged by the limitations of finite basis sets. Two particularly intertwined phenomena arise from this limitation: the Basis Set Superposition Error (BSSE) and the Basis Set Incompleteness Error (BSIE). While both stem from the use of incomplete basis sets, they manifest differently and require distinct correction strategies. BSSE is an artificial lowering of energy that specifically plagues interaction energy calculations between molecular fragments. It occurs because fragments in a molecular complex "borrow" basis functions from neighboring fragments, achieving a lower energy state not through genuine interaction but through an expanded, unbalanced basis set representation [5] [19]. Conversely, BSIE is the general error in any electronic structure calculation arising from the failure to describe the wavefunction completely, most notably the electron-electron cusp condition [20]. This guide provides a comprehensive comparison of the primary methodological approaches—the a posteriori counterpoise (CP) correction and the a priori Chemical Hamiltonian Approach (CHA)—developed to overcome these challenges, evaluating their theoretical foundations, accuracy, and practical implementation.
Theoretical Foundations: BSSE and BSIE
The Nature of Basis Set Superposition Error (BSSE)
BSSE is a notorious problem in the study of weakly bonded molecular complexes. In a typical interaction energy calculation, the energy of the complex (supermolecule) is compared to the sum of the energies of the isolated monomers. The error arises because the supermolecule calculation benefits from a more complete basis set—the combined basis sets of all monomers—while the monomer calculations are performed with their individual, smaller basis sets. This leads to a spurious overestimation of the binding energy [19]. The error is formally defined in the Boys-Bernardi counterpoise (CP) scheme, where the interaction energy is corrected as: ΔEABCP = EAB(AB) - EA(AB) - EB(AB) Here, the notation EA(AB) signifies the energy of monomer A calculated with the full basis set of the complex AB [19]. This correction aims to provide a balanced treatment by using the same basis set for all components of the energy difference.
The Basis Set Incompleteness Error (BSIE) and the CBS Limit
BSIE is a more fundamental error present in all electronic structure calculations that employ a finite basis set. The exact wavefunction satisfies the electron cusp condition, a sharp feature in the wavefunction at electron coalescence points due to the divergence of the Coulomb potential [20]. Smooth Gaussian-type orbitals (GTOs), the standard in quantum chemistry, fail to capture this cusp, necessitating very large basis sets for accurate results. The Complete Basis Set (CBS) limit is the theoretical result obtained with an infinitely large basis set. Since this is computationally unattainable, extrapolation techniques are employed. A common two-point extrapolation scheme by Helgaker et al. uses the formula: E∞ ≈ (EXX3 - EX-1(X-1)3) / (X3 - (X-1)3) where X is the cardinal number of the basis set (e.g., 2, 3, 4 for double-, triple-, zeta sets) and EX is the energy computed with that basis set [21]. The slow convergence of energy with basis set size makes BSIE a significant source of uncertainty.
The following conceptual diagram illustrates the relationship and primary correction pathways for these two errors:
Methodological Approaches and Comparative Accuracy
A Posteriori Correction: The Counterpoise (CP) Method
The CP method, introduced by Boys and Bernardi, is the most widespread technique for BSSE correction [19] [22]. It is an a posteriori correction, meaning it is applied after the individual energy calculations. The core concept of the CP method is the use of ghost atoms—atoms with basis functions but no nuclear charge or electrons. These are placed at the positions of the partner monomer to recreate the full dimer basis set during monomer energy calculations [5] [22]. The following workflow details the steps for a CP-corrected interaction energy calculation:
A Priori Elimination: The Chemical Hamiltonian Approach (CHA)
In contrast to CP, the Chemical Hamiltonian Approach (CHA) is an a priori method designed to be inherently free of BSSE. It achieves this by modifying the Hamiltonian itself to prevent the unphysical delocalization of electrons that causes BSSE. A key theoretical distinction is that CHA employs a non-Hermitian Fock matrix. This is justified because BSSE is not a physical phenomenon and does not correspond to an observable with a Hermitian operator [19]. The energy in the CHA method is ultimately computed as the expectation value of the standard, Hermitian Hamiltonian using the BSSE-free CHA wavefunctions, ensuring a real-valued energy [19].
Quantitative Comparison of Methodological Accuracy
The performance of CP, CHA, and other methods has been rigorously tested on model systems. The table below summarizes key experimental data for the water dimer, a benchmark system for hydrogen bonding studies, obtained with a 4-31G basis set [19].
Table 1: Comparison of SCF Interaction Energies (kcal/mol) for Water Dimer at O–O Distance = 2.8 Å
| Method | Interaction Energy | BSSE Treatment | Includes Physical CT? |
|---|---|---|---|
| Uncorrected SCF | -5.66 | None | Yes |
| CP-Corrected | -4.81 | A Posteriori | Yes |
| CHA/F | -4.86 | A Priori | Yes |
| SCF-MI | ~ -3.5 | A Priori (Incorrect) | No |
The data reveals crucial insights. The CP and CHA methods yield remarkably similar interaction energies (-4.81 and -4.86 kcal/mol, respectively), suggesting both are valid and effective approaches for BSSE correction. In contrast, the SCF-MI method, which constrains molecular orbitals to individual monomers, significantly overcorrects and gives a much weaker binding energy. This is because it not only removes BSSE but also eliminates the physical charge-transfer (CT) effects between monomers, which are crucial for a proper description of hydrogen bonding [19]. This underscores a critical point: a valid BSSE correction must remove the artificial stabilization without sacrificing the physically real intermolecular interactions.
Advanced Protocols and Modern Solutions
Protocol for Counterpoise Correction with Ghost Atoms
Implementing the CP correction requires careful setup. Here is a detailed protocol based on standard quantum chemistry software practices:
- Geometry Input: The input deck must specify the atomic coordinates of the complex and, crucially, the positions of the ghost atoms. These are the coordinates of the partner monomer(s).
- Ghost Atom Specification: Ghost atoms are defined with zero nuclear charge and zero mass. They are typically designated in the
$moleculesection with the atomic symbolGhor by prefixing an actual atomic symbol with@(e.g.,@Ofor an oxygen ghost atom) [22]. - Basis Set Assignment: The basis set for the ghost atoms must be explicitly assigned. This can be done in a
$basissection withBASIS = MIXED, where each atom (real and ghost) is assigned its original basis set [5] [22]. When using the@symbol, the ghost atom automatically inherits the basis set of the corresponding real atom. - Energy Calculations: Perform three separate single-point energy calculations: (i) the full complex
E_AB(AB), (ii) monomer A in the presence of ghost atoms for BE_A(AB), and (iii) monomer B in the presence of ghost atoms for AE_B(AB). - Energy Combination: Use the CP formula to compute the final, corrected interaction energy.
Tackling BSIE: CBS Extrapolation and Explicitly Correlated Methods
While CP and CHA address BSSE, tackling BSIE requires separate strategies to approach the CBS limit.
CBS Extrapolation and Error Estimation: As noted in Section 2.2, CBS extrapolation is a common technique. A significant challenge is estimating the uncertainty of the extrapolated value. A modern approach involves using ensemble random walks to simulate all possible extrapolation outcomes that could be obtained with larger, unavailable basis sets. By analyzing the statistical distribution of these outcomes, one can assign reliable confidence intervals to the CBS limit estimate, providing a system-specific and non-empirical uncertainty quantification [21].
Explicitly Correlated Methods (F12): These methods, such as F12 and Transcorrelated (TC) theory, directly incorporate the electron-electron distance (r12) into the wavefunction Ansatz. This explicitly satisfies the cusp condition, dramatically accelerating convergence to the CBS limit. The primary advantage is achieving chemical accuracy with vastly smaller basis sets [20]. For example, the TC approach performs a similarity transformation on the Hamiltonian, transferring correlation from the wavefunction into the Hamiltonian itself. This results in a more compact ground state, which is particularly beneficial for quantum computing algorithms, as it reduces the required number of qubits and circuit depth [20]. The trade-off is that the TC Hamiltonian becomes non-Hermitian, requiring specialized algorithms like variational Quantum Imaginary Time Evolution (VarQITE) for its solution on quantum hardware [20].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Computational Tools and Concepts for BSSE and BSIE Research
| Tool/Concept | Function/Role | Example/Note |
|---|---|---|
| Ghost Atoms (Gh) | Basis functions without nuclear charge, used in CP corrections to model the partner monomer's basis set. | Defined with zero mass and charge [5]. |
| Correlation-Consistent Basis Sets (cc-pVXZ) | A systematic series of basis sets designed for controlled convergence to the CBS limit. | X = D, T, Q, 5... (cardinal numbers) [21]. |
| Counterpoise (CP) Protocol | The standard a posteriori procedure for calculating BSSE-corrected interaction energies. | Requires multiple single-point calculations with ghost atoms [22]. |
| Chemical Hamiltonian Approach (CHA) | An a priori method that defines a BSSE-free Hamiltonian. | Results in a non-Hermitian Fock matrix [19]. |
| Explicitly Correlated Methods (F12/R12) | Introduces explicit electron-distance dependence to rapidly converge energies to the CBS limit. | Reduces BSIE, allowing smaller basis sets for high accuracy [20]. |
| Complete Basis Set (CBS) Extrapolation | A mathematical technique to estimate the CBS limit results from calculations with finite basis sets. | Often uses a Helgaker-style (X-3) extrapolation scheme [21]. |
The comparative analysis of CP and CHA reveals a nuanced landscape. The CP method is a robust, widely implemented, and empirically successful a posteriori correction. Its main strength is its conceptual simplicity and direct approach to balancing the basis sets. Criticisms that it may overcorrect are noted in the literature, but comparative studies show the degree of undercorrection assumed by some critics is "quite unrealistic," and its results are in close agreement with the a priori CHA method [19]. CHA's principal advantage is its theoretical elegance, removing BSSE at its source without the need for multiple corrective calculations. The requirement for non-Hermitian machinery is a theoretical strength, not a weakness, given the non-observable nature of BSSE [19].
For the practicing computational chemist, the choice of method is often practical. The CP method is integrated into most major quantum chemistry software packages (e.g., ADF, Q-Chem) and is the de facto standard [5] [22]. CHA, while theoretically sound, has seen less widespread adoption in mainstream computational workflows. Ultimately, for the demanding task of predicting reliable intermolecular interaction energies—be it in drug design, materials science, or catalysis—a dual strategy is essential. One must simultaneously correct for BSSE using a validated method like CP or CHA, and mitigate BSIE by employing large basis sets with CBS extrapolation or, preferably, explicitly correlated F12-type methods. They are indeed "two sides of the same coin," both rooted in basis set incompleteness, and both must be addressed to uncover the true chemical physics of molecular interactions.
Implementing BSSE Corrections: A Step-by-Step Guide to CP and CHA in Practice
In the computational study of weak molecular interactions, such as those critical in drug design and materials science, achieving accurate interaction energies is paramount. A significant challenge in these calculations is the Basis Set Superposition Error (BSSE). This error arises when calculating a molecular dimer (A-B); the basis functions on fragment A artificially help to lower the energy of fragment B, and vice versa. This results in an interaction energy that is biased towards dimer formation due purely to basis set effects, not genuine physical interaction [23]. The Counterpoise (CP) correction, specifically the Boys-Bernardi Counterpoise Correction (BB-CP), is a systematic procedure designed to correct for this deficiency [23]. It estimates what the energies of the isolated monomers would be if they were calculated with the full dimer basis set, thereby providing a less biased interaction energy. The accuracy of such quantum-mechanical benchmarks is crucial, as errors even as small as 1 kcal/mol can lead to erroneous conclusions in fields like drug design [24].
Methodology and Theoretical Foundation
The Boys-Bernardi Formulation
The core of the CP correction lies in the original Boys and Bernardi formula. The BSSE-corrected interaction energy, ΔE, between fragments A and B is given by:
[ \Delta E = E^{AB}{AB}(AB) - E^{A}{A}(A) - E^{B}{B}(B) - \left[E^{AB}{A}(AB) - E^{AB}{A}(A) + E^{AB}{B}(AB) - E^{AB}_{B}(B)\right] ]
In this notation, (E_{X}^{Y} (Z)) represents the energy of fragment X calculated at the optimized geometry of fragment Y with the basis set of fragment Z [23]. The terms in the square brackets constitute the BSSE correction itself. The equation can be interpreted as follows: the first three terms represent the uncorrected interaction energy calculated in their own bases, while the terms within the bracket adjust for the artificial stabilization of the monomers by the partner's basis functions.
The Role of 'Ghost Atoms'
The term "ghost atom" is central to the practical implementation of the CP correction. A ghost atom is an atom that provides its basis functions for the quantum chemical calculation but contributes no electrons or nuclear charge to the Hamiltonian [23]. This allows for a computation where a monomer's energy is calculated in the presence of the complete dimer basis set, but without the physical presence of the other monomer's atoms, thus isolating the effect of the basis set. In computational chemistry packages like ORCA, this is typically achieved by placing a colon (":") after the atomic symbol in the coordinate input. For example, an oxygen ghost atom would be specified as O: [23]. This simple syntax instructs the software to include the basis set for that atom as a placeholder, enabling the crucial single-point energy calculations needed for the BB-CP formula.
Workflow for Counterpoise Correction
Implementing a full CP correction requires a series of coordinated quantum chemical calculations. The following workflow, depicted in the diagram below, outlines the essential steps for a dimer A-B.
Diagram 1: CP Correction Workflow
The workflow consists of the following key steps:
- Geometry Optimizations: First, the geometries of the dimer (A-B) and the individual monomers (A and B) are optimized independently with their own basis sets. This yields the energies (E{AB}^{AB}(AB)), (E{A}^{A}(A)), and (E_{B}^{B}(B)) [23].
- Single-Point at Dimer Geometry: The optimized dimer structure is taken, and one fragment (e.g., A) is deleted. A single-point energy calculation is run on the remaining fragment (B) at this geometry with its original basis set. This is repeated for the other fragment, giving (E{A}^{AB}(A)) and (E{B}^{AB}(B)) [23].
- Single-Point with Ghost Basis: This is the crucial "ghost atom" step. Using the optimized dimer geometry again, the energy of monomer A is calculated, but this time with the full dimer basis set. This is achieved by specifying the atoms of monomer B as ghost atoms. The process is repeated for monomer B, yielding (E{A}^{AB}(AB)) and (E{B}^{AB}(AB)) [23].
- Energy Computation: All the calculated energies are inserted into the Boys-Bernardi formula to obtain the BSSE-corrected interaction energy.
Experimental Protocols and Data Presentation
A Practical Example: The Water Dimer
To illustrate the CP correction, consider a study of the water dimer at the MP2/cc-pVTZ level of theory. The following table summarizes the energies required and the resulting corrected interaction energy [23].
Table 1: CP-Corrected Energy Calculation for Water Dimer [23]
| Energy Component | Description | Energy (a.u.) |
|---|---|---|
| (E^{AB}_{AB}(AB)) | Energy of the optimized dimer | -152.646980 |
| (E^{A}_{A}(A)) | Energy of optimized monomer A | -76.318651 |
| (E^{B}_{B}(B)) | Energy of optimized monomer B | -76.318651 |
| (E^{AB}_{A}(AB)) | Energy of monomer A with ghost basis of B | -76.320799 |
| (E^{AB}_{A}(A)) | Energy of monomer A at dimer geometry | -76.318635 |
| (E^{AB}_{B}(AB)) | Energy of monomer B with ghost basis of A | -76.319100 |
| (E^{AB}_{B}(B)) | Energy of monomer B at dimer geometry | -76.318605 |
From these values, the interaction energies are calculated as:
- Uncorrected Dimer Energy: (\Delta E{dim.} = E^{AB}{AB}(AB) - E^{A}{A}(A) - E^{B}{B}(B) = -0.009677) a.u. (-6.07 kcal/mol)
- BSSE Correction: (\Delta E{BB-CP} = [E^{AB}{A}(AB) - E^{AB}{A}(A)] + [E^{AB}{B}(AB) - E^{AB}_{B}(B)] = 0.002659) a.u. (1.67 kcal/mol)
- Corrected Interaction Energy: (\Delta E{dim., corr.} = \Delta E{dim.} - \Delta E_{BB-CP} = -0.007018) a.u. (-4.40 kcal/mol)
This example shows that the BSSE can be a significant fraction of the total interaction energy, in this case over 25%, underscoring the importance of the CP correction for quantitative accuracy.
Benchmarking Performance in Complex Systems
The critical role of high-accuracy benchmarks like those corrected for BSSE is highlighted by the "QUantum Interacting Dimer" (QUID) framework, designed for ligand-pocket interactions in drug design [24]. QUID uses a "platinum standard" for interaction energies, established by achieving tight agreement (within 0.5 kcal/mol) between two entirely different high-level quantum methods: Linearized Coupled Cluster Singles and Doubles with perturbative Triples (LNO-CCSD(T)) and Fixed-Node Diffusion Monte Carlo (FN-DMC) [24]. This approach minimizes uncertainty and provides a robust benchmark for assessing faster, more approximate methods.
Table 2: Performance of Computational Methods on the QUID Benchmark [24]
| Method Type | Example Method(s) | Typical Performance on QUID | Key Limitations |
|---|---|---|---|
| Gold/Platinum Standard | LNO-CCSD(T), FN-DMC | Reference (Error ~0.5 kcal/mol) | Computationally prohibitive for large systems. |
| Density Functional Theory | PBE0+MBD, B3LYP-D3(BJ) | Accurate for energy, but atomic forces may differ. | Quality heavily depends on the chosen functional and dispersion correction. |
| Semiempirical Methods | GFNn-xTB | Requires improvement for non-equilibrium geometries. | Often fail to capture the full complexity of NCIs. |
| Empirical Force Fields | Standard MMFFs | Require improvement for NCIs and transferability. | Use effective pairwise approximations; lack explicit polarization. |
The benchmark analysis reveals that while several dispersion-inclusive density functional approximations can provide accurate energy predictions, their descriptions of atomic forces (e.g., van der Waals forces) can differ in both magnitude and orientation [24]. Furthermore, semiempirical methods and empirical force fields generally require further improvement to reliably capture non-covalent interactions, especially for out-of-equilibrium geometries sampled during binding events [24].
Implementation in Software and Advanced Applications
Software-Specific Syntax
The implementation of ghost atoms and CP corrections varies slightly between computational chemistry packages. The following table serves as a quick-reference "toolkit" for researchers.
Table 3: Research Reagent Solutions for CP Corrections
| Tool / Concept | Software | Implementation Example | Function |
|---|---|---|---|
| Ghost Atom | ORCA | O : 5.752050 6.489306 5.407671 |
Includes basis set for oxygen at given coordinates, without electrons/nucleus [23]. |
| Ghost Fragment | ORCA | GhostFrags {1} end |
Defines an entire molecular fragment as ghost atoms [23]. |
| CP Correction | Psi4 | bsse_type='cp' |
Automates the computation of CP-corrected interaction energies [25]. |
| CP Geometry Opt. | ORCA | BSSEOptimization.cmp |
Compound script for geometry optimizations with CP correction [23]. |
| Geom. CP (gCP) | ORCA | N/A (Automatic) | Adds semi-empirical BSSE correction to HF/DFT energies; corrects intramolecular BSSE [23]. |
Beyond Single-Point Energies: Geometries and Dynamics
The CP correction's application has expanded beyond single-point energy calculations. For instance, ORCA now supports geometry optimizations with analytic counterpoise corrections, allowing for the determination of accurate non-covalent complex geometries, not just energies [23]. This is enabled by specialized compound scripts rather than a simple !Opt keyword.
Another advanced approach is the Geometrical Counterpoise correction (gCP). This method adds a semi-empirical correction, (E{\text{gCP}}), directly to the HF or DFT energy: (E{\text{total}} = E{\text{HF/DFT}} + E{\text{gCP}}) [23]. Parametrized to approximate the Boys-Bernardi CP correction, gCP has the distinct advantage of also correcting for intramolecular BSSE and is computationally inexpensive as it requires no additional electronic structure calculations [23].
The Counterpoise correction remains a foundational technique for achieving accurate interaction energies in computational chemistry. Its methodology, centered on the use of "ghost atoms" to eliminate basis set superposition error, provides a systematic workflow that is implemented in major quantum chemistry software. While the full Boys-Bernardi CP correction is computationally demanding, its importance is underscored by high-level benchmarks like QUID, which show that even modern DFT methods can have subtle inaccuracies that rigorous corrections help to uncover. The development of advanced extensions, such as CP-corrected geometry optimizations and semi-empirical gCP schemes, continues to enhance its utility, making it an indispensable tool for researchers and drug development professionals who require high fidelity in modeling non-covalent interactions.
The accurate computation of interaction energies in molecular complexes and clusters is a cornerstone of computational chemistry, with critical applications in drug development for predicting ligand-receptor binding and solvation effects. A significant challenge in these calculations is the Basis Set Superposition Error (BSSE), an artifact of using incomplete basis sets. BSSE leads to an artificial overestimation of binding energy because the basis functions of one molecule (or fragment) can be used to lower the energy of its interacting partner, a phenomenon known as basis set mixing. Two philosophically distinct strategies have been developed to address this issue: the a posteriori Counterpoise (CP) correction and the a priori Chemical Hamiltonian Approach (CHA).
The Counterpoise (CP) correction, introduced by Boys and Bernardi, is a widely accepted a posteriori method. It corrects the interaction energy after the fact by recalculating the energy of each monomer using the entire basis set of the supermolecule [26]. In contrast, the Chemical Hamiltonian Approach (CHA) seeks to prevent basis set mixing from the outset by constructing a Hamiltonian that is fundamentally free from the effects of BSSE. While the CP method is empirical and has been extensively documented, the CHA is a more theoretical construct designed to avoid the error inherently. This guide provides an objective comparison of these methodologies, focusing on their theoretical foundations, practical performance, and applicability in modern computational research, particularly for pharmaceutical development.
Theoretical Foundations and Methodologies
The Counterpoise (CP) Correction Method
The CP correction is a pragmatic approach applied after the computation of the complex's energy. The standard supermolecule interaction energy (ΔEINT) is calculated as the difference between the energy of the complex and the sum of the energies of the isolated monomers, each in their own basis set [26]. The BSSE arises because the monomers in the complex benefit from a larger, combined basis set (χ<sub>M1,M2,…,MN</sub>), compared to their isolated state basis sets (χ<sub>Mi</sub>).
The CP correction accounts for this by re-defining the interaction energy (ΔECP-INT) such that the energy of each isolated monomer is also computed using the full, supersystem basis set [26]. This eliminates the energy advantage gained from basis set mixing, as shown in the equation below:
ΔECP-INT = EχM1,M2,…,MNM1M2…MN - Σi=1N EχM1,M2,…,MNMi [citation:]
The CP scheme has been validated across numerous systems, from dimers to many-body clusters. Studies have shown that CP-corrected Hartree-Fock interaction energies become largely basis-set independent, even with moderate-sized basis sets like cc-pVDZ, facilitating more reliable predictions without the need for prohibitively large basis sets [26].
The Chemical Hamiltonian Approach (CHA)
The Chemical Hamiltonian Approach represents a more fundamental solution. Instead of correcting the energy after the calculation, CHA designs the Hamiltonian operator itself to be independent of the basis sets of other fragments. The core idea is to formulate a Hamiltonian that describes the true electronic structure of the individual molecules without the artificial stabilization afforded by the "borrowing" of basis functions from neighbors. This a priori prevention aims to yield interaction energies that are inherently BSSE-free from the start of the calculation, without requiring any additional correction steps.
While the theoretical elegance of CHA is appealing, its practical implementation in mainstream quantum chemistry software and its application to large, complex systems like drug-like molecules are less common compared to the widely implemented CP correction. The subsequent sections will compare these methods based on available experimental data, much of which benchmarks against CP-corrected results.
Comparative Performance Analysis
Performance in Intermolecular Complexes and Clusters
The performance of BSSE correction methods is critical in many-body systems, which are more representative of real-world environments in materials science and drug discovery than simple dimers. Research has demonstrated that the conventional CP correction effectively recovers BSSE in many-body clusters of organic compounds. A study on the 3B-69 dataset (consisting of 69 trimers from organic crystal structures) and the MBC-36 dataset (containing clusters of 2, 4, 8, and 16 molecules from benzene, aspirin, and oxalyl dihydrazide polymorphs) confirmed that CP-corrected HF interaction energies were basis-set independent across a series of Dunning's basis sets (cc-pVXZ and aug-cc-pVXZ, X = D, T) [26].
Table 1: Performance of CP Correction in Many-Body Clusters [26]
| Dataset | Cluster Types | Key Finding | Basis Sets Used |
|---|---|---|---|
| 3B-69 | Trimers (69 systems) | CP-corrected interaction energies are basis-set independent. | cc-pVDZ, cc-pVTZ, aug-cc-pVDZ, aug-cc-pVTZ |
| MBC-36 | Dimers, Tetramers, Octamers, 16-mers | A cut-off radius of ~10 Å is sufficient to recover BSSE effects in crystalline environments. | cc-pVDZ, cc-pVTZ, aug-cc-pVDZ, aug-cc-pVTZ |
Furthermore, the study found that using a relatively small basis set like cc-pVDZ with CP correction showed excellent performance in predicting HF interaction energies, offering a cost-effective strategy for large clusters [26]. The local nature of BSSE was also established, with a cut-off radius of 10 Å being sufficient to fully recover these effects in crystal structures [26].
Performance in Geometry Optimizations
The influence of BSSE correction on molecular geometries is another crucial metric. A benchmarking study on 21 van der Waals dimers provides insights into how the CP correction affects optimized structures. The study used CCSD(T)/CBS as a reference and compared geometries optimized with methods like MP2 and CCSD with various basis sets.
Table 2: Impact of CP Correction on Geometry Optimizations of Van der Waals Dimers [27]
| Basis Set Size | Effect of Counterpoise (CP) Correction | Recommendation |
|---|---|---|
| Double-Zeta (e.g., aug-cc-pVDZ) | Tends to degrade the quality of optimized geometries. | Avoid CP correction during geometry optimization with small basis sets. |
| Triple-Zeta and Larger (e.g., aug-cc-pVTZ) | Has a larger effect, generally improving convergence towards the CBS limit. | Using CP correction with larger basis sets is beneficial for accurate geometries. |
The study concluded that the frozen core approximation induces only very small geometric changes, while the CP correction has a more significant impact [27]. This highlights a nuanced picture: while CP is invaluable for accurate energy calculations, its application in geometry optimization must be considered with care, particularly with smaller basis sets.
A separate B3LYP study on hydrated complexes of [K(H2O)n]+ and [Na(H2O)n]+ (n=1–6) further underscores the importance of method selection. The research found that basis sets like 6-31G* were inadequate for accurate hydration energies, while larger basis sets like 6-31+G* and 6-31++G were necessary. CP-corrected geometry optimizations were crucial for predicting properties like hydration distances and energies that agreed with experimental results [28].
Experimental Protocols and Workflows
Standard Protocol for Counterpoise-Corrected Interaction Energy Calculation
For a molecular cluster, the protocol for calculating the CP-corrected interaction energy at the Hartree-Fock level is as follows [26]:
- Cluster Geometry Extraction: Obtain the geometry of the N-body cluster from a reliable source, such as an experimental crystal structure (e.g., from the Cambridge Structural Database) or a pre-optimized structure.
- Supermolecule Energy Calculation: Perform a single-point energy calculation on the entire cluster (the supermolecule) using a chosen quantum chemistry method (e.g., HF) and basis set (
χ<sub>M1,M2,...,MN</sub>). This yieldsE<sub>χ<sub>M1,M2,…,MN</sub></sub><sup>M1M2…MN</sup>. - Monomer Energy in Full Basis Set: For each monomer
iin the cluster, perform a single-point energy calculation using the same method and the full basis set of the cluster (χ<sub>M1,M2,...,MN</sub>). The monomer geometry must be kept identical to its geometry within the cluster. This yieldsE<sub>χ<sub>M1,M2,…,MN</sub></sub><sup>Mi</sup>for each monomer. - Energy Calculation: Compute the CP-corrected interaction energy (ΔECP-INT) using the equation provided in Section 2.1.
The following workflow diagram illustrates this process:
Workflow for BSSE-Corrected Geometry Optimization
For systems where geometry is sensitive to BSSE, such as hydrated metal ions, a CP-corrected optimization can be performed [28]:
- Initial Guess: Provide an initial guess for the molecular geometry.
- Energy & Gradient with CP: For the current geometry, compute the energy and the molecular gradient (first derivative of energy with respect to nuclear coordinates) using the Counterpoise method. This means the gradient calculation also accounts for BSSE.
- Geometry Update: The optimizer uses the CP-corrected gradient to determine a new, lower-energy geometry.
- Convergence Check: Steps 2 and 3 are repeated until geometric parameters (e.g., bond distances, angles) and the energy meet specified convergence criteria.
The Scientist's Toolkit: Essential Research Reagents
The table below lists key computational "reagents" and methodologies used in the featured studies for BSSE research.
Table 3: Key Research Reagents and Methods in BSSE Studies
| Item / Method | Function in BSSE Research | Example from Literature |
|---|---|---|
| Dunning's Correlation-Consistent Basis Sets | A family of basis sets (e.g., cc-pVXZ, aug-cc-pVXZ) designed for systematic convergence to the Complete Basis Set (CBS) limit, allowing study of BSSE dependence. | Used to demonstrate basis-set independence of CP-corrected energies in the 3B-69 and MBC-36 datasets [26]. |
| Benchmark Datasets (3B-69, MBC-36) | Curated collections of molecular clusters extracted from crystal structures, providing standardized systems for testing and benchmarking quantum chemical methods. | Used to evaluate CP correction performance in many-body systems beyond dimers [26]. |
| Supermolecule Approach | The foundational computational method for calculating interaction energies by treating a molecular cluster as a single quantum mechanical entity. | The basis for both uncorrected and CP-corrected interaction energy calculations [26]. |
| CCSD(T)/CBS | A high-accuracy coupled-cluster method considered the "gold standard" for providing reference interaction energies and geometries against which other methods are benchmarked. | Used as a reference to assess the quality of CP-corrected geometries of van der Waals dimers [27]. |
| Counterpoise (CP) Algorithm | The standard computational procedure for correcting BSSE by recalculating monomer energies in the full supersystem basis set. | Implemented in geometry optimizations of [K(H₂O)ₙ]⁺ and [Na(H₂O)ₙ]⁺ clusters to study hydration energies and structures [28]. |
The search for an optimal strategy to address Basis Set Superposition Error remains a active area of research in computational chemistry. The evidence indicates that the Counterpoise correction is a robust, well-validated, and practical method for calculating accurate interaction energies, particularly for many-body clusters relevant to pharmaceutical and materials science [26]. Its ability to produce basis-set-independent results with moderately sized basis sets offers a compelling balance of accuracy and computational cost.
The Chemical Hamiltonian Approach, while theoretically elegant as an a priori method, currently lacks the same extensive benchmarking and widespread implementation in mainstream studies. The available literature heavily focuses on validating and understanding the CP scheme across diverse systems. Future research may provide more direct comparative data between CHA and CP, especially for large, complex biological systems. For now, the CP correction, applied with an understanding of its limitations—such as its variable effect on geometry optimization with different basis sets [27]—represents the most empirically grounded and widely used tool for correcting BSSE in drug development research.
Accurate calculation of binding energies is foundational to understanding weak, non-covalent interactions in host-guest complexes and dimers. These interactions, essential in drug design, supramolecular chemistry, and materials science, often feature binding energies that are small differences between large molecular energies. This makes them highly susceptible to numerical error, particularly from the Basis Set Superposition Error (BSSE). BSSE is an artificial lowering of energy that occurs when finite basis sets are used, as atoms from one molecule借用 the basis functions of another to describe their electron density more completely. Two prominent methodological approaches have been developed to correct for this error: the Counterpoise (CP) Correction method, proposed by Boys and Bernardi, and the Chemical Hamiltonian Approach (CHA). This guide provides an objective comparison of their application, performance, and accuracy within modern computational workflows, providing researchers with the data needed to select an appropriate correction strategy.
Theoretical Frameworks and Experimental Protocols
The Counterpoise Correction Protocol
The Counterpoise (CP) method is a post-calculation correction applied to the interaction energy. The standard protocol for a host-guest complex, as implemented in studies of calix[4]arene complexes and chloride anion hosts, involves several key steps [29] [30]:
- Geometry Optimization: The geometry of the complex (AB), the isolated host (A), and the isolated guest (B) are optimized at a chosen level of theory (e.g., DFT with a functional like B3LYP-D3 and a basis set like 6-311+G(d,p) or def2-TZVP).
- Single-Point Energy Calculations: The energies of all three species are calculated using the geometry of the complex.
- ( E{AB}(AB) ): Energy of the complex with its own basis set.
- ( E{A}(A) ): Energy of the isolated host with its own basis set.
- ( E_{B}(B) ): Energy of the isolated guest with its own basis set.
- Ghost Orbital Calculations: The energies of the host and guest are recalculated using the full basis set of the complex.
- ( E{A}(AB) ): Energy of the host in the geometry of the complex, with the guest present as a "ghost" (its nucleus is removed but its basis functions remain).
- ( E{B}(AB) ): Energy of the guest in the geometry of the complex, with the host present as a "ghost."
- BSSE-Corrected Interaction Energy: The CP-corrected interaction energy (( \Delta E_{CP} )) is computed as:
- ( \Delta E{CP} = E{AB}(AB) - [E{A}(AB) + E{B}(AB)] ) The uncorrected interaction energy is ( \Delta E{Uncorrected} = E{AB}(AB) - [E{A}(A) + E{B}(B)] ).
This protocol is widely supported in major computational chemistry software packages like Gaussian [29] [30].
The Chemical Hamiltonian Approach Protocol
The Chemical Hamiltonian Approach (CHA) is an a priori* method that aims to eliminate BSSE from the outset by redefining the Hamiltonian of the system. The general workflow is:
- Hamiltonian Formulation: The standard electronic Hamiltonian is partitioned to explicitly exclude the overlap between basis functions on different monomers. This creates a BSSE-free "chemical Hamiltonian."
- Wavefunction Calculation: The Schrödinger equation is solved for the complex using this modified Hamiltonian. The resulting interaction energy is inherently corrected for BSSE without the need for additional ghost orbital calculations.
- Property Calculation: Molecular properties are derived directly from the wavefunction obtained with the chemical Hamiltonian.
While theoretically elegant, practical implementations of CHA are less common in mainstream quantum chemistry software compared to the Counterpoise method.
Standardized Workflow for Accuracy Comparison
To objectively compare the accuracy of CP and CHA, a controlled computational experiment should be designed. The following workflow, derived from benchmark creation and host-guest studies, outlines this process [29] [30] [31].
Performance Comparison: Counterpoise Correction vs. Chemical Hamiltonian Approach
A direct, quantitative comparison of CP and CHA in the context of the provided search results is challenging. The current literature, as evidenced by studies on host-guest complexes and benchmark dataset creation, heavily favors the use of Counterpoise correction, with no specific experimental data for CHA found in the searched sources. The following table summarizes the objective findings based on the available data.
Table 1: Comparative Analysis of Counterpoise and Chemical Hamiltonian Approach Based on Searched Literature
| Feature | Counterpoise (CP) Correction | Chemical Hamiltonian Approach (CHA) |
|---|---|---|
| Theoretical Principle | A posteriori empirical correction of the interaction energy [30]. | A priori reformulation of the system's Hamiltonian to exclude BSSE. |
| Implementation Prevalence | Extremely Common. Directly implemented and used in major software (Gaussian) for studying host-guest systems [29] [30]. | Not Reported. The searched literature did not mention its use in recent experimental studies. |
| Reported Quantitative Performance | Considered a standard, necessary step for reliable interaction energies in calix[4]arene/solvent complexes and chloride anion host-guest systems [29] [30]. | No comparative performance data was available in the searched sources. |
| Computational Cost | Requires additional "ghost" energy calculations, increasing the number of single-point computations [30]. | Theoretical cost is lower per calculation, but overall workflow efficiency is unreported. |
| Key Applications in Literature | Correction of interaction energies in calix[4]arene/solvent complexes [29] and chlorine isotope effect studies [30]. | Not mentioned in the context of the host-guest and dimer studies found in the search results. |
The search results indicate that the Counterpoise method is the de facto standard for BSSE correction in practical research settings involving host-guest complexes and dimers. For instance, a 2025 study on calix[4]arene inclusion complexes explicitly states, "the counterpoise approach is used to modify the interaction energies in order to eliminate the basis set superposition error (BSSE)" [29]. Similarly, a 2021 study on isotopic consequences of host-guest interactions notes the application of BSSE correction for gas-phase complexes, though it also highlights that such correction is not available for continuum solvent models in some computational protocols [30].
The Scientist's Toolkit: Essential Research Reagents and Solutions
The following table details key computational tools and methodologies identified in the literature for accurately modeling and correcting binding energies in weak interactions.
Table 2: Key Research Reagent Solutions for Energy Correction Studies
| Tool/Solution | Function in Research | Example Use Case |
|---|---|---|
| Counterpoise Correction | Empirically corrects for BSSE in interaction energy calculations by using "ghost" orbitals [30]. | Essential for calculating accurate binding energies in calix[4]arene/organic solvent inclusion complexes [29]. |
| Implicit Solvent Models (e.g., CPCM, SMD) | Models the effect of a solvent environment using a continuous dielectric medium, crucial for simulating realistic conditions [29]. | Used to calculate interaction energies of CX[4]/solvent systems in a solvent phase, complementing gas-phase studies [29]. |
| SAPT (Symmetry-Adapted Perturbation Theory) | Decomposes interaction energy into physically meaningful components (electrostatics, exchange, induction, dispersion), providing deep insight into binding forces [31]. | Used to create the Splinter dataset for benchmarking protein-ligand interaction energies and training machine learning models [31]. |
| Benchmark Datasets (e.g., Splinter) | Provides high-quality reference data (e.g., from SAPT calculations) for training, testing, and validating new computational methods and force fields [31]. | Contains ~1.6 million dimer configurations with SAPT0-calculated energies to improve prediction of protein-ligand interactions [31]. |
The current computational chemistry landscape for studying weak interactions is heavily oriented toward the Counterpoise correction method. The search results confirm its entrenched position as a necessary and standard tool for obtaining reliable binding energies in host-guest complexes and dimers [29] [30]. In contrast, the Chemical Hamiltonian Approach, while a significant theoretical concept, is not reflected in contemporary experimental protocols within the scope of the searched literature.
The field is progressing toward the creation of large, high-quality benchmark datasets like Splinter, which use advanced methods like SAPT to provide reference data [31]. The future of accurate binding energy calculation likely lies in the development and refinement of machine learning models and force fields trained on such data, offering speed without sacrificing quantum mechanical accuracy. For the practicing researcher today, the Counterpoise method remains the essential, empirically validated choice for BSSE correction, while CHA's practical utility and performance in modern applications require further independent investigation.
Addressing Intramolecular BSSE in Covalent Bond Breaking and Conformational Energy Calculations
The basis set superposition error (BSSE) is traditionally recognized as a critical issue in calculating intermolecular non-covalent interactions, where the finite basis set of one molecule artificially lowers the energy of another by providing additional basis functions. However, a more insidious form of this error—intramolecular BSSE—permeates all electronic structure calculations involving covalent bond breaking, conformational changes, and transition state analysis [13]. This error arises when one part of a molecule "borrows" basis functions from another region within the same molecule, leading to unbalanced descriptions of different molecular structures and systematically biased relative energies [13] [1].
Within the context of methodological comparisons, two primary approaches have emerged to correct BSSE: the counterpoise (CP) correction and the Chemical Hamiltonian Approach (CHA). While the CP method calculates and subtracts the BSSE a posteriori by introducing "ghost orbitals," the CHA prevents basis set mixing a priori by modifying the Hamiltonian itself [1]. Understanding the relative performance of these approaches is particularly crucial for researchers investigating chemical reactions, drug design, and materials science, where accurate energy differences dictate predictive reliability. This guide provides an objective comparison of these methodologies, supported by experimental data and practical protocols for their implementation.
Theoretical Framework: CP vs. CHA
Fundamental Principles and Computational Mechanisms
The Counterpoise (CP) Correction, introduced by Boys and Bernardi, is the most widely employed BSSE correction scheme [7] [25]. It operates on a simple principle: the energy of each molecular fragment (monomer) should be evaluated using the same complete basis set as the supermolecule (e.g., the complex or the entire molecule in a specific conformation). For a dimer system A-B, the CP-corrected interaction energy is calculated as:
[ \Delta E{CP} = E{AB}^{AB}(AB) - E{AB}^{AB}(A) - E{AB}^{AB}(B) ]
where the notation (E{P}^{B}(F)) denotes the energy of fragment F calculated with the basis set of system P, at the geometry of system G [7] [1]. The terms (E{AB}^{AB}(A)) and (E_{AB}^{AB}(B)) represent the energies of the isolated monomers computed with the full dimer basis set, often implemented using "ghost atoms" that provide basis functions without atomic nuclei or electrons.
In contrast, the Chemical Hamiltonian Approach (CHA) represents a more fundamental solution by modifying the quantum chemical Hamiltonian to eliminate terms that lead to BSSE [1]. Instead of an a posteriori correction, the CHA constructs a Hamiltonian that is manifestly free from BSSE by "projecting out" the components that allow one fragment to artificially lower the energy of another through basis set borrowing [1]. Although conceptually elegant and theoretically sound, the CHA formalism is less developed for complex chemical reactions compared to the CP method [7].
Key Differentiating Factors
Table 1: Fundamental Comparison of CP and CHA Approaches
| Feature | Counterpoise Correction | Chemical Hamiltonian Approach |
|---|---|---|
| Philosophy | A posteriori correction | A priori prevention |
| Implementation | "Ghost orbital" calculations | Modified Hamiltonian |
| Theoretical Status | Well-established for intermolecular complexes | Less developed for reaction pathways [7] |
| Computational Cost | Additional single-point calculations | Modified integral evaluation |
| Fragment Definition | Requires explicit fragment assignment | Requires explicit fragment assignment |
| Basis Set Dependence | Error diminishes with larger basis sets [1] | Error diminishes with larger basis sets [1] |
Comparative Accuracy in Chemical Reactions
The PH₃ + H → PH₂ + H₂ Model Reaction
A seminal study by László et al. systematically investigated BSSE corrections for the model reaction PH₃ + H → PH₂ + H₂ using various levels of theory (HF, MP2, MP4, QCISD, QCISD(T)) and basis sets [7]. This work highlights the critical challenge of applying BSSE corrections to covalent bond-breaking and forming processes, particularly in the transition state region.
The researchers identified two significant problems when applying CP corrections to reaction barriers:
- Mathematical Ambiguity: For asymmetric reactions, the barrier height becomes ill-defined because the CP correction computed with respect to reactants differs from that computed with respect to products [7].
- Physical Assignment Problem: Near the transition state, the transferred atom (hydrogen in this case) cannot be legitimately assigned to either fragment, necessitating a three-fragment treatment for which the standard two-fragment CP method is inadequate [7].
To address the three-fragment problem, generalized CP schemes have been proposed. The Turi-Dannenberg scheme computes each monomer's energy in the full supermolecule basis, while the White-Davidson/Valiron-Mayer approach uses a hierarchical scheme with separate CP corrections for 1-body, 2-body, etc., energy components [7].
Table 2: BSSE Correction Effects on Reaction Energy and Barrier (PH₃ + H → PH₂ + H₂) at MP2/6-311+G* Level [7]*
| Method | Reaction Energy (kcal/mol) | Barrier Height (kcal/mol) |
|---|---|---|
| Uncorrected | -14.2 | 4.8 |
| Standard CP (2-fragment) | -13.5 | 5.8 |
| Generalized CP (3-fragment) | -13.1 | 6.5 |
The data reveals that BSSE corrections significantly impact both reaction energies and barrier heights, with the magnitude of correction depending on the specific CP scheme employed. The lack of a "perfect and practicable solution" underscores the need for careful methodological choices based on the specific chemical system [7].
Performance in Conformational Energy Calculations: The Proton Affinity Benchmark
Beyond reaction pathways, intramolecular BSSE substantially affects the calculation of conformational energies and molecular properties like proton affinity (PA) and gas-phase basicity (GPB) [13]. Systematic studies on hydrocarbons demonstrate that BSSE is not confined to large molecules or weak interactions but permeates all electronic structure calculations with limited basis sets.
The intramolecular BSSE in these systems arises because the description of the protonation site is artificially improved in the conjugate acid by "borrowing" basis functions from other parts of the molecule, similar to the intermolecular case [13]. This leads to systematically biased proton affinities when using smaller basis sets, with the error direction and magnitude depending on the molecular structure and basis set quality.
For drug development professionals, these findings are particularly relevant when computing pKa values, binding affinities, or conformational energy landscapes, where even small energy errors (1-2 kcal/mol) can dramatically impact predictive accuracy.
Experimental Protocols and Implementation
Standard Counterpoise Correction Protocol
For implementing CP corrections in intermolecular interactions or intramolecular fragment analyses, follow this standardized protocol:
- Geometry Optimization: Optimize the geometry of the complete system (supermolecule) at your chosen level of theory without BSSE correction.
- Single-Point Energy Calculation: Perform a single-point energy calculation on the optimized geometry with the CP correction specified. In many quantum chemistry packages, this involves:
- Defining molecular fragments in the input file
- Including keywords such as
counterpoise=N(in Gaussian) or using thebsse_typeparameter (in Psi4) [25]
- Fragment Energy Calculation: The program automatically computes:
- The total supermolecule energy, (E{AB}^{AB}(AB))
- Fragment A energy in the full basis, (E{AB}^{AB}(A))
- Fragment B energy in the full basis, (E_{AB}^{AB}(B))
- Energy Calculation: The CP-corrected interaction energy is: [ \Delta E{CP} = E{AB}^{AB}(AB) - E{AB}^{AB}(A) - E{AB}^{AB}(B) ]
For intramolecular BSSE assessment in conformational analysis, apply the same protocol by treating different conformational states as distinct "supermolecules" and dividing the molecule into logically defined fragments.
Workflow for Transition State BSSE Assessment
Assessing BSSE in transition states requires special considerations due to the fragment assignment problem [7]:
- Multiple Fragment Definitions: Calculate CP corrections using different fragment assignments (reactant-like and product-like partitions).
- Three-Fragment Treatment: For reactions involving atom transfer, implement a three-fragment scheme where the transferred atom constitutes a separate fragment.
- Barrier Consistency Check: Compute the forward and backward reaction barriers with both fragment definitions; significant discrepancies indicate heightened BSSE sensitivity.
- Basis Set Convergence: Repeat the analysis with increasingly larger basis sets to monitor BSSE reduction.
Figure 1: Workflow for BSSE assessment in transition states, highlighting the need for multiple fragment definitions.
Research Reagent Solutions: Computational Tools for BSSE Correction
Table 3: Essential Software and Computational Tools for BSSE Research
| Tool Name | Type/Function | BSSE Capabilities |
|---|---|---|
| Psi4 | Quantum Chemistry Package | Comprehensive BSSE correction via bsse_type keyword (CP, NoCP, VMFC) [25] |
| Gaussian | Quantum Chemistry Package | Counterpoise correction with fragment-based input |
| libint | Integral Library | Provides one- and two-electron integrals for new DFT implementations [32] |
| PYSEQM | Semiempirical QM Package | Python-based SEQM methods compatible with ML parameterization [33] |
| HIPNN+SEQM | Machine Learning Model | Dynamically parameterized Hamiltonians to improve accuracy [33] |
Emerging Approaches and Future Directions
Machine Learning-Enhanced Quantum Chemistry
Recent advances in machine learning (ML) offer promising alternatives to traditional BSSE correction methods. The HIPNN+SEQM framework, for instance, replaces static semiempirical parameters with dynamically generated values inferred from the local chemical environment [33]. This approach maintains the quantum mechanical structure while significantly improving accuracy, achieving improved transferability to larger systems with reduced BSSE sensitivity.
Another approach, OrbNet, utilizes symmetry-adapted atomic orbital features from semiempirical calculations to achieve high learning efficiency while reducing computational cost [33]. These ML-integrated methods represent a paradigm shift where physical knowledge is embedded directly into scalable, accurate models that inherently mitigate basis set deficiencies.
Methodological Recommendations for Different Use Cases
Based on the comparative analysis:
- For drug discovery professionals calculating conformational energies or binding affinities, always test the sensitivity of your results to BSSE by comparing CP-corrected and uncorrected values with at least a moderate basis set (e.g., 6-311+G).
- For reaction mechanism studies involving covalent bond breaking/forming, implement the three-fragment CP scheme for transition states and report both reactant- and product-corrected barriers as an uncertainty measure.
- For high-accuracy benchmarking, combine large basis sets with CP corrections, as BSSE diminishes more rapidly with basis set size than the inherent basis set incompleteness error [1].
- For large systems where CP corrections become prohibitive, consider ML-enhanced quantum methods like HIPNN+SEQM that show promising transferability while maintaining physical interpretability [33].
Figure 2: Decision framework for selecting appropriate BSSE correction strategies based on system type.
Computational chemistry provides a suite of electronic structure methods, each with distinct strengths, limitations, and domains of applicability. For researchers in drug development and materials science, selecting the appropriate method is critical for obtaining reliable predictions of molecular properties, reaction energies, and non-covalent interactions. This guide objectively compares the performance of three foundational families of methods—Hartree-Fock (HF), Density Functional Theory (DFT), and post-Hartree-Fock (post-HF) theories—framed within research assessing the accuracy of counterpoise corrections and the chemical Hamiltonian approach for addressing basis set superposition errors. Understanding the theoretical underpinnings, systematic errors, and computational costs of these methods is a prerequisite for evaluating such specialized correction techniques.
Fundamental Methodologies
Hartree-Fock (HF) Theory: HF is a wavefunction-based method that serves as the starting point for most advanced quantum chemical calculations. It treats electrons as moving independently in an average field created by other electrons but explicitly enforces the Pauli exclusion principle through the antisymmetry of its wavefunction, written as a Slater determinant [34]. Its major limitation is the complete neglect of electron correlation, the energy associated with correlated electron motion beyond this mean-field approximation.
Density Functional Theory (DFT): DFT is an alternative formulation that uses the electron density, rather than the many-electron wavefunction, as the fundamental variable [35]. While exact in principle, the unknown exchange-correlation functional must be approximated. The accuracy of a DFT calculation depends almost entirely on the quality of this functional, leading to a vast "Jacob's Ladder" of approximations [35].
Post-Hartree-Fock Methods: This category includes methods designed to recover the electron correlation missing in HF. They range from moderately expensive Møller-Plesset perturbation theory (e.g., MP2) to highly accurate but computationally intensive approaches like Coupled-Cluster (CC) theory, particularly CCSD(T), which is often regarded as the "gold standard" of quantum chemistry for its high accuracy [36].
The Challenge of Basis Set Superposition Error (BSSE)
When calculating interaction energies between molecular fragments, a significant error can arise from the use of finite basis sets. The Basis Set Superposition Error (BSSE) is an artificial lowering of the energy of a dimer complex because each fragment can "borrow" basis functions from the other, improving its own description compared to an isolated calculation [36]. Two primary strategies exist to mitigate BSSE:
- Counterpoise (CP) Correction: A widely used technique where the energy of each isolated fragment is recalculated using the full dimer basis set [36].
- Chemical Hamiltonian Approach (CHA): An alternative method that projects out the BSSE from the Hamiltonian at the outset of the calculation.
Benchmarking the performance of HF, DFT, and post-HF methods often involves evaluating their susceptibility to BSSE and the efficacy of these correction schemes.
Comparative Performance Analysis
Accuracy Across Chemical Systems
The performance of electronic structure methods is highly system-dependent. The following table summarizes their typical performance for different chemical properties and systems.
Table 1: Performance of Electronic Structure Methods Across Different Chemical Problems
| Chemical System / Property | HF Performance | DFT Performance | Post-HF Performance | Key Evidence |
|---|---|---|---|---|
| Zwitterions & Charge-Transfer Systems | Excellent for structure-property correlation; handles localization well [34]. | Poor with global hybrids; improved with range-separated hybrids (RSH) due to delocalization error [34] [35]. | Excellent; CCSD, CASSCF, etc., match HF and experiment for zwitterions [34]. | HF reproduced exp. dipole moment (~10.33D) of pyridinium benzimidazolate; DFT (B3LYP, etc.) showed significant deviations [34]. |
| Non-Covalent Interactions (NCIs) | Poor; lacks dispersion interactions, leading to severe underbinding. | Variable; semi-local functionals fail. Requires empirical dispersion corrections (e.g., -D4) or vdW-inclusive functionals [36]. | Excellent; CCSD(T) is the benchmark for NCIs, inherently capturing dispersion [31] [36]. | MLIPs trained on CCSD(T) data achieve chemical accuracy for vdW-dominated systems and protein-ligand interactions [31] [36]. |
| Transition Metal Complexes & Strong Correlation | Poor; cannot handle multi-reference character. | Variable to Poor; standard functionals fail for open-shell systems, charge transfer, and strongly correlated bonds [37]. | Excellent when multi-reference methods (e.g., CASSCF) are used [37]. | Large Wavefunction Models (LWMs) show promise in capturing strong static correlation where DFT fails [37]. |
| Geometries & Bond Lengths | Fair; tends to overestimate bond lengths due to lack of correlation [35]. | Good to Excellent; GGA and hybrid functionals are generally reliable for geometry optimization [35] [38]. | Excellent; especially with CCSD(T) and large basis sets [36]. | MIC DFT-D computations accurately reproduce high-quality X-ray structures [38]. |
| Reaction Energetics & Thermochemistry | Poor; no correlation energy, unreliable barriers and reaction energies. | Variable; meta-GGAs and hybrids like BMK, M06-2X can be good, but results are functional-dependent [34]. | Excellent; CCSD(T) near the CBS limit is the reference standard [36]. |
Computational Cost and Scalability
A critical practical consideration is the computational resource requirement, which determines the feasible system size.
Table 2: Computational Scaling and Practical Considerations
| Method | Formal Scaling | Typical Maximum System Size (Atoms) | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Hartree-Fock (HF) | ( O(N^4) ) | Hundreds | Simple, no functional choice, good for localized electrons [34]. | Neglects electron correlation, poor for NCIs and thermochemistry. |
| Density Functional Theory (DFT) | ( O(N^3) ) to ( O(N^4) ) | Thousands (pure GGA) / Hundreds (hybrid) | Excellent cost-accuracy trade-off [39]. | Functional-dependent results; delocalization/self-interaction errors [39] [35]. |
| MP2 | ( O(N^5) ) | Tens to Hundreds | Includes electron correlation, captures dispersion. | Susceptible to BSSE; fails for metallic/multi-reference systems. |
| CCSD(T) | ( O(N^7) ) | Dozens (canonical) / Hundreds (local) | "Gold standard" for accuracy [36]. | Prohibitively expensive for large systems; the ( O(N^7) ) scaling is a major bottleneck [37] [36]. |
Figure 1: A hierarchical overview of common electronic structure methods, their key features, and limitations. The choice of method involves a trade-off between computational cost and physical accuracy.
Experimental Protocols for Method Benchmarking
Protocol 1: Benchmarking Zwitterionic Molecular Properties
This protocol is based on a study investigating the performance of HF and DFT on pyridinium benzimidazolate zwitterions [34].
- Objective: To assess the ability of various methods to reproduce experimental molecular structures and dipole moments of zwitterions.
- Software Used: Gaussian 09 [34].
- Methods Compared: A wide range including HF, multiple DFT functionals (B3LYP, CAM-B3LYP, BMK, M06-2X, etc.), and post-HF methods (MP2, CASSCF, CCSD, QCISD) [34].
- Procedure:
- Geometry Optimization: All molecular structures are fully optimized without symmetry constraints using each method.
- Frequency Calculation: Vibrational frequency calculations are performed to confirm the structures are true local minima (no imaginary frequencies).
- Property Calculation: Dipole moments are computed from the optimized electron density.
- Validation: Computed structures (e.g., twist angles between aryl rings) and dipole moments are compared against high-resolution experimental X-ray crystallography data.
- Key Findings: HF and high-level post-HF methods (CCSD, CASSCF) reproduced experimental dipole moments and structures with high fidelity, while many global hybrid DFT functionals performed poorly. This was attributed to HF's advantage in handling localized charge states, unlike DFT's tendency for spurious delocalization [34].
Protocol 2: Δ-Learning for CCSD(T)-Accuracy Potentials
This protocol describes a machine-learning workflow to achieve CCSD(T) accuracy for periodic systems, a common challenge in material science and drug development [36].
- Objective: To create machine-learning interatomic potentials (MLIPs) that match CCSD(T) accuracy for systems dominated by van der Waals interactions, at a fraction of the computational cost.
- Software Used: MOLPRO 2024.1 for quantum chemical calculations [36].
- Reference Method: PNO-LCCSD(T)-F12/haTZ (a highly accurate, locally approximated coupled-cluster method with explicit correlation) [36].
- Procedure:
- Baseline Calculation: A low-cost, dispersion-corrected tight-binding (DFT) calculation is performed.
- Δ-Learning: An MLIP is trained not on the total CCSD(T) energy, but on the energy difference ((\Delta)) between the target CCSD(T) energy and the baseline DFT energy. This is more efficient than learning the total energy from scratch.
- Training Set: The model is trained on a diverse set of molecular fragments and vdW-bound multimers to ensure transferability of dispersion interactions.
- BSSE Handling: The F12 explicit correlation method dramatically reduces basis-set incompleteness error, thereby minimizing BSSE without the need for an explicit counterpoise correction [36].
- Key Findings: The resulting (\Delta)-learned potential achieved root-mean-square energy errors below 0.4 meV/atom and accurately reproduced interaction energies, bond lengths, and vibrational frequencies at CCSD(T) quality for covalent organic frameworks (COFs) and other systems [36].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Software, Methods, and Basis Sets for Electronic Structure Calculations
| Tool Name | Type | Primary Function | Relevance to BSSE/CHA Research |
|---|---|---|---|
| Gaussian 09 | Software Suite | General-purpose quantum chemistry package for HF, DFT, and post-HF calculations [34]. | Used for foundational studies comparing method performance on molecular properties, providing data for BSSE analysis [34]. |
| MOLPRO | Software Suite | High-accuracy quantum chemistry package specializing in correlated wavefunction methods [36]. | Used to generate benchmark CCSD(T) reference data with low BSSE (via F12 correction) for training ML models [36]. |
| PNO-LCCSD(T)-F12 | Quantum Method | Local coupled-cluster, the gold standard for correlated energies in large systems [36]. | Serves as the near-exact reference for benchmarking lower-level methods and correction schemes; its inherent low BSSE is a key advantage [36]. |
| Counterpoise (CP) Correction | Computational Protocol | A posteriori correction for BSSE in interaction energy calculations [36]. | The standard against which the performance of other approaches (like CHA) is measured. |
| aug-cc-pVXZ (e.g., haTZ) | Basis Set Family | Correlation-consistent basis sets augmented with diffuse functions [36]. | Reduces BSSE by providing a more complete description of the electron density, especially in regions critical for weak interactions. Essential for high-accuracy work. |
| Machine-Learning Interatomic Potentials (MLIPs) | AI Model | Fast, empirical potentials trained on QM data to achieve QM accuracy [36]. | (\Delta)-learning on CCSD(T) data provides a path to bypass the cost and BSSE issues of direct DFT or HF calculations for large systems [36]. |
| Range-Separated Hybrid (RSH) Functionals | DFT Functional | Class of DFT functionals that mix HF and DFT exchange in a distance-dependent manner [35]. | Mitigates delocalization error in DFT, improving performance for charge-transfer and zwitterionic systems where BSSE and self-interaction are problematic [34] [35]. |
Emerging Trends and Future Directions
The field is rapidly evolving with the integration of artificial intelligence and novel computational frameworks. Machine learning is now being used not just for interatomic potentials but also to learn and correct for fundamental errors in DFT approximations directly [39]. Furthermore, Large Wavefunction Models (LWMs) represent a paradigm shift. These are foundation neural-network wavefunctions optimized by Variational Monte Carlo (VMC) that directly approximate the many-electron wavefunction, offering a path to achieve gold-standard accuracy without the prohibitive ( O(N^7) ) scaling of traditional CCSD(T) [37]. These approaches promise to generate large-scale, high-accuracy ab-initio datasets, which will be invaluable for refining existing methods and correction protocols like counterpoise and CHA.
Troubleshooting BSSE Corrections: Overcoming Inconsistencies and Optimization Strategies
In the pursuit of accurate quantum chemical predictions, researchers have long grappled with the basis set superposition error (BSSE), an artifact arising from the use of incomplete basis sets in electronic structure calculations. This error manifests as an overestimation of binding energies in weakly bonded complexes because monomers in a dimer system can "use" the basis functions of their partner, a capability unavailable in isolated monomer calculations [7]. For decades, the counterpoise (CP) correction method, introduced by Boys and Bernardi, has served as the predominant solution to estimate and eliminate BSSE, becoming virtually ubiquitous in studies of intermolecular interactions [7] [40].
The standard CP approach computes interaction energy by recalculating monomer energies using the entire basis set of the complex, effectively isolating the pure interaction energy from basis set artifacts [7]. However, this conventional wisdom faces mounting theoretical and practical challenges when extended beyond simple dimer systems, particularly for mapping potential energy surfaces of chemical reactions. This analysis examines the fundamental inconsistencies and dangers inherent in applying CP corrections across reactive potential energy surfaces, contrasting its performance with emerging alternatives that offer more physically consistent pathways to accuracy.
Theoretical Foundations: CP Methodology and Its Limitations
The Fundamental CP Correction Scheme
The CP method operates on a straightforward principle: to correct for the artificial stabilization caused by BSSE, one must compute the energy of each monomer (fragment) using not only its own basis functions but also those of the interacting partner. For a dimer system AB, the CP-corrected interaction energy is calculated as:
[ \Delta E{CP} = E{AB}(AB) - [EA(AB) + EB(AB)] ]
Where $E{AB}(AB)$ represents the energy of the dimer in the full basis set, while $EA(AB)$ and $E_B(AB)$ represent the energies of monomers A and B computed in the full dimer basis set [7]. This scheme aims to provide a consistent basis set for comparing dimer and monomer energies, thereby eliminating the BSSE advantage enjoyed by the dimer system.
Inherent Limitations and Conceptual Challenges
Despite its widespread adoption, the CP method suffers from significant theoretical shortcomings:
Directional Dependence Problem: For asymmetric reactions, the CP correction becomes ill-defined at the transition state because the correction differs depending on whether fragments are defined with respect to reactants or products [7]. This creates an ambiguous barrier height, fundamentally undermining the method's reliability for kinetic studies.
Fragment Assignment Challenge: Near the transition state of bimolecular reactions, the transferred atom or group cannot be legitimately assigned to either reactant, necessitating a three-fragment treatment for which the standard two-fragment CP scheme is inadequate [7].
Questionable Foundation: Empirical evidence suggests that, contrary to the fundamental hypothesis behind CP correction, standard quantum chemistry basis sets may actually be "biased toward the atom" with larger basis set errors for dimers than monomers. In such cases, applying CP correction increases the imbalance rather than mitigating it [40].
Comparative Analysis: Counterpoise Correction vs. Chemical Hamiltonian Approach
Performance Evaluation on Model Systems
Studies on prototype systems reveal critical limitations of CP corrections. Research on the $\text{PH}3 + \text{H} \rightarrow \text{PH}2 + \text{H}_2$ reaction demonstrated significant conceptual problems when applying CP corrections to potential energy surfaces, manifesting as non-negligible changes to barrier heights that cast doubt on reaction mechanism predictions [7].
Table 1: Performance Comparison of BSSE Correction Methods for Reaction Energy Surfaces
| Method | Theoretical Basis | Transition State Treatment | Fragment Assignment | Barrier Height Consistency |
|---|---|---|---|---|
| Counterpoise Correction | Corrects monomer energies in dimer basis | Directionally dependent; ill-defined | Problematic for transferred atoms | Inconsistent for asymmetric reactions |
| Chemical Hamiltonian Approach (CHA) | Modifies Hamiltonian to exclude BSSE terms | Physically consistent framework | Naturally handles multiple fragments | Theoretically more consistent |
| Uncorrected Calculations | No BSSE correction | Naturally continuous | Straightforward but inaccurate | Consistent but quantitatively erroneous |
Perhaps more damning are investigations of simple systems like $\text{Be}_2$, where researchers could compare CP results against sufficiently accurate benchmark numbers (close to basis set limit and full CI). These studies found that "counterpoise corrected bond energies then deviate more from the basis set limit numbers than uncorrected bond energies" [40]. This conclusion held at both the Hartree-Fock level and, more strongly, at correlated levels like CCSD(T) and full CI.
The Chemical Hamiltonian Alternative
The Chemical Hamiltonian Approach (CHA) represents a fundamentally different strategy that addresses BSSE at its source. Instead of applying posteriori corrections to energies, CHA identifies and excludes the specific terms in the Hamiltonian responsible for BSSE [7]. Experience shows that CHA and CP methods typically yield similar results for intermolecular interaction energies when significant BSSE contaminates uncorrected results. However, a critical advantage of CHA is its more robust theoretical foundation, though its practical implementation remains limited as "the CHA formalism has not yet been developed for reactions" [7].
Experimental Protocols and Methodologies
Standard CP Implementation Protocol
The typical workflow for applying CP correction to potential energy surfaces involves:
Geometry Optimization: Locate stationary points (reactants, products, complexes, transition states) on the uncorrected potential energy surface.
Single-Point CP Calculations: For each geometry, compute:
- The total energy of the complete system
- Individual fragment energies using the entire supermolecule basis set
Energy Correction: Apply the CP correction formula at each point: [ E{CP} = E{AB}^{AB} - [E{A}^{AB} + E{B}^{AB}] ]
Surface Mapping: Construct the corrected potential energy surface using CP-corrected energies.
This protocol faces particular difficulties at transition states, where fragment definition becomes ambiguous, especially for asymmetric reactions [7].
Advanced Multi-Fragment CP Schemes
To address the limitations in transition state regions, researchers have proposed generalized CP schemes:
- Turi-Dannenberg Scheme: Each monomer energy is computed in the entire supermolecule basis [7].
- White-Davidson/Valiron-Mayer Scheme: A hierarchical approach decomposing total energy into 1-body, 2-body contributions with separate CP corrections for each component [7].
These methods remain computationally demanding and lack universal adoption, highlighting the practical challenges in achieving consistent BSSE correction across full potential energy surfaces.
Diagram 1: Workflow and failure points for CP correction on reaction energy surfaces. The process highlights critical ambiguities at transition states that lead to inconsistent predictions.
Emerging Alternatives and Modern Approaches
Explicitly Correlated Methods
Beyond CHA, explicitly correlated methods offer promising alternatives to traditional CP correction:
Transcorrelated (TC) Approach: Incorporates correlated Ansätze directly into the Hamiltonian via similarity transformation, effectively building electron correlation into the Hamiltonian itself [20]. This method reduces the need for large basis set expansions by directly incorporating the electronic cusp condition, significantly reducing BSSE at its source rather than applying posteriori corrections.
Canonical Transcorrelated F12 (CT-F12): Uses unitary operators in similarity transformation, maintaining Hermitian Hamiltonians while incorporating correlation effects, though it requires truncation of the Baker-Campbell-Hausdorff expansion [20].
Deep Learning Hamiltonian Prediction
Recent advances in machine learning offer a fundamentally different approach to electronic structure calculation:
NextHAM Framework: A neural E(3)-symmetry and expressive correction method that predicts electronic-structure Hamiltonians directly from atomic configurations [41]. This approach circumvents the self-consistent field procedure entirely, potentially avoiding BSSE issues associated with traditional quantum chemistry calculations while offering dramatic computational efficiency improvements.
Materials-HAM-SOC Dataset: A benchmark containing 17,000 material structures spanning 68 elements, enabling training and evaluation of generalizable Hamiltonian prediction models without traditional BSSE complications [41].
Quantum Computing Approaches
Quantum computational chemistry represents another frontier for overcoming traditional limitations:
Variational Quantum Eigensolver (VQE) with TC Integration: Combining transcorrelated methods with variational quantum algorithms reduces both qubit requirements and circuit depths, enabling more accurate calculations on near-term quantum hardware [20].
Sample-based Quantum Diagonalization: Leverages quantum computers as sampling engines to define subspaces where Hamiltonians are diagonalized classically, offering promising alternatives to conventional approaches [42].
Table 2: Modern Computational Approaches for BSSE Mitigation
| Method | Mechanism | BSSE Handling | Implementation Status | Key Advantages |
|---|---|---|---|---|
| Explicitly Correlated (F12) Methods | Incorporates explicit electron distance dependence | Reduces BSSE through better wave function description | Established in specialized codes | Faster basis set convergence |
| Transcorrelated (TC) Approach | Similarity transformation of Hamiltonian | Builds correlation directly into Hamiltonian | Quantum computing implementations | Reduced qubit requirements, shallower circuits |
| Deep Learning Hamiltonians | Neural network prediction from atomic structures | Bypasses traditional SCF procedure | Emerging research framework | DFT-level precision with superior computational efficiency |
| Quantum Algorithm Integration | Quantum-classical hybrid approaches | Inherits advantages of incorporated methods | Early experimental stage | Potential for quantum advantage |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Computational Tools and Methods for Electronic Structure Studies
| Tool/Resource | Function | Application Context |
|---|---|---|
| GAUSSIAN-94 Program [7] | Electronic structure package | Traditional CP correction implementation |
| 6-311+G Basis Set [7] | Moderate-sized Gaussian-type orbital basis | Balanced accuracy/efficiency for reaction studies |
| Materials-HAM-SOC Dataset [41] | Broad-coverage materials benchmark | Training/evaluating machine learning models |
| Transcorrelated Ansatz [20] | Explicitly correlated wave function | Hamiltonian transformation for reduced BSSE |
| Variational Quantum Eigensolver [20] | Hybrid quantum-classical algorithm | Ground state energy calculation on quantum hardware |
The theoretical inconsistencies and empirical evidence demonstrate significant dangers in applying standard CP corrections across full potential energy surfaces, particularly for chemical reactions. The directional dependence problem, fragment assignment ambiguity, and fundamental questions about its validity challenge the routine application of CP methodology for reaction kinetics studies [7] [40].
While CP correction remains useful for simple intermolecular complexes, researchers studying reaction mechanisms should approach it with caution and consider emerging alternatives. The Chemical Hamiltonian Approach offers a more fundamental solution but requires further development for reaction applications [7]. Meanwhile, explicitly correlated methods, deep learning Hamiltonian prediction, and quantum computational approaches represent promising pathways toward more consistent and accurate electronic structure predictions without the inherent contradictions of surface-wide CP correction [41] [20].
The field increasingly recognizes that the pursuit of chemical accuracy requires moving beyond corrective patches to fundamentally better theoretical frameworks that avoid the problems altogether rather than attempting to correct them posteriori.
In quantum chemical simulations, the selection of an appropriate basis set—a collection of mathematical functions used to represent molecular orbitals—represents a fundamental compromise between computational cost and accuracy. This balance is particularly crucial in drug discovery, where researchers must model complex molecular interactions with sufficient precision while working within practical computational constraints. The inevitable tradeoff between runtime and accuracy determines the applicability of quantum chemical calculations across many domains, as the size of addressable systems is often limited by computational resources rather than intrinsic simulation error [6]. Basis set selection directly impacts two significant error sources: Basis Set Incompleteness Error (BSIE), which arises from an insufficient number of basis functions, and Basis Set Superposition Error (BSSE), an artificial overestimation of interaction energies when fragments "borrow" adjacent basis functions from each other [6]. This guide objectively compares contemporary basis set strategies and error correction methodologies, providing researchers with evidence-based recommendations for selecting optimal approaches specific to their computational challenges.
Types of Basis Set Errors
Basis Set Incompleteness Error (BSIE) originates from using a finite number of basis functions to represent molecular orbitals, preventing the wavefunction from reaching the complete basis set (CBS) limit. As the basis set expands toward this limit, the total energy typically decreases, and molecular properties become more accurate [43]. BSIE is especially problematic for weak non-covalent interactions like van der Waals forces and hydrogen bonding, where even small energy errors (1 kcal/mol) can lead to erroneous conclusions about relative binding affinities in drug design [44].
Basis Set Superposition Error (BSSE) represents an artificial lowering of energy in molecular complexes compared to separated fragments. This occurs because basis functions on one fragment provide additional variational flexibility for electrons on adjacent fragments. In interaction energy calculations, BSSE leads to overestimated binding energies, as fragments temporarily access each other's basis functions [14]. The magnitude of BSSE depends on both basis set size and the nature of molecular interactions, with smaller basis sets typically exhibiting larger errors [14].
Error Correction Approaches
Two primary methodologies address BSSE: the Counterpoise (CP) Correction and the Chemical Hamiltonian Approach (CHA). The CP method, proposed by Boys and Bernardi, calculates BSSE by performing "ghost orbital" calculations where monomers are evaluated using the full dimer basis set [14]. The CP-corrected interaction energy is obtained by comparing these ghost orbital calculations with standard monomer calculations. While CP and CHA tend to yield similar results despite conceptual differences, CP has gained wider adoption due to its conceptual simplicity and ease of implementation [14]. Recent research indicates that CP corrections remain valuable even with high-level methods, though their importance diminishes with larger, more complete basis sets [43].
Table 1: Comparison of Error Correction Methods for Basis Set Artifacts
| Method | Theoretical Basis | Implementation Complexity | Systematic Error | Computational Cost |
|---|---|---|---|---|
| Counterpoise (CP) | Ghost orbital calculations | Low | Tends to underbind | Moderate (requires additional monomer calculations) |
| Chemical Hamiltonian Approach (CHA) | Projection operators to eliminate BSSE | High | More variable | Lower (no additional calculations) |
| Implicit (Large Basis Sets) | Basis set completeness reduces errors | None | Diminishes with basis set size | High (increased basis functions) |
Experimental Protocols for Basis Set Evaluation
Benchmarking Strategies and Dataset Selection
Robust evaluation of basis set performance requires standardized benchmarking on chemically diverse datasets with high-level reference data. The GMTKN55 main-group thermochemistry benchmark set has emerged as a standard for quantifying DFT method accuracy, encompassing various reaction types including basic properties, isomerization, barrier heights, and non-covalent interactions [6]. For non-covalent interactions specifically, specialized benchmarks like the A24 dataset (24 noncovalently bound dimers), S66, and the more recent QUID (QUantum Interacting Dimer) framework provide rigorous testing grounds [44] [43]. These datasets span diverse interaction types including hydrogen bonding, dispersion, and mixed-character complexes, enabling comprehensive assessment of basis set performance across different chemical environments.
Computational Methodology Specifications
High-quality benchmarking requires careful attention to computational parameters. Studies should employ sufficiently dense integration grids—for example, a (99,590) grid with "robust" pruning using the Stratmann–Scuseria–Frisch quadrature scheme [6]. Tight SCF convergence criteria and integral tolerances (e.g., 10⁻¹⁴) help minimize numerical errors unrelated to basis set choice [6]. For geometry optimizations, which are particularly sensitive to BSSE, protocols should specify whether optimizations were performed on CP-corrected or uncorrected potential energy surfaces, as this significantly impacts resulting molecular structures [14]. When comparing against reference methods like CCSD(T), researchers should employ explicitly correlated F12 corrections with large basis sets (e.g., heavy-aug-cc-pVTZ) to minimize residual BSIEs in the reference values themselves [36].
Basis Set Selection and Evaluation Workflow
Comparative Performance Analysis of Basis Set Strategies
Conventional Basis Sets and Their Performance Characteristics
Traditional basis sets follow systematic construction principles, with the Dunning cc-pVXZ (correlation-consistent polarized valence X-zeta) family representing a gold standard for method benchmarking. These basis sets demonstrate predictable convergence toward the complete basis set limit with increasing cardinal number X (D, T, Q, 5), but require careful handling of BSSE [43]. For noncovalent interactions, the addition of diffuse functions (aug-cc-pVXZ) proves essential for accurate description of electron density tails in van der Waals complexes [43]. Popular alternatives like Pople-style basis sets (e.g., 6-311+G(d)) offer reasonable accuracy at lower computational cost but exhibit less systematic convergence [14].
Recent benchmarking reveals that conventional double-ζ basis sets like def2-SVP and 6-31G(d) suffer from substantial pathologies, including poor electron density description and overestimated interaction energies [6]. One comprehensive study noted that increasing from double-ζ (def2-SVP) to triple-ζ (def2-TZVP) basis sets caused calculation runtimes to increase more than five-fold, creating significant practical constraints for drug discovery applications on large or conformationally flexible systems [6].
Table 2: Performance Comparison of Basis Sets Across Methodologies (Weighted Total Mean Absolute Deviations - WTMAD2)
| Methodology | def2-QZVP | vDZP | def2-SVP | 6-31G(d) | pcseg-1 |
|---|---|---|---|---|---|
| B97-D3BJ | 8.42 | 9.56 | 12.84 | 14.92 | 11.63 |
| r2SCAN-D4 | 7.45 | 8.34 | 10.27 | 11.85 | 9.76 |
| B3LYP-D4 | 6.42 | 7.87 | 9.43 | 10.68 | 8.91 |
| M06-2X | 5.68 | 7.13 | 8.76 | 9.94 | 8.12 |
Data derived from GMTKN55 benchmarks as reported in [6]. Lower values indicate better performance.
Emerging Efficient Basis Sets and Composite Approaches
The recently developed vDZP basis set represents a significant advancement in balancing cost and accuracy. Specifically designed to minimize BSSE almost to triple-ζ levels, vDZP employs effective core potentials and deeply contracted valence basis functions optimized on molecular systems [6]. Remarkably, vDZP demonstrates broad applicability across multiple density functionals without method-specific reparameterization. In comprehensive evaluations, vDZP-based methods substantially outperform conventional double-ζ basis sets while maintaining similar computational costs [6].
The frozen natural orbital (FNO) approach provides another strategy for cost reduction, particularly valuable for quantum computing applications like quantum phase estimation (QPE). By truncating parts of the virtual orbital space while starting from a large parent basis set, FNOs capture dynamical correlation effects with significantly reduced computational resources—achieving up to 80% reduction in the Hamiltonian 1-norm (λ) and 55% reduction in orbital count while preserving accuracy [45].
Table 3: Key Research Reagents and Computational Resources for Basis Set Studies
| Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| GMTKN55 Database | Benchmark Dataset | Comprehensive main-group thermochemistry reference | Method validation and comparison |
| QUID Framework | Benchmark Dataset | 170 non-covalent dimers modeling ligand-pocket interactions | Biological binding energy assessment |
| vDZP Basis Set | Basis Set | Double-zeta basis with minimal BSSE | Cost-effective DFT calculations |
| cc-pVXZ Family | Basis Set | Systematic correlation-consistent basis sets | High-accuracy reference calculations |
| Counterpoise Correction | Algorithm | BSSE correction via ghost orbitals | Accurate interaction energy computation |
| Frozen Natural Orbitals | Method | Virtual space truncation | Resource reduction in high-level methods |
Integrated Recommendations for Different Research Scenarios
Basis Set Selection Guidelines
For routine DFT calculations on medium-sized systems (50-200 atoms), the vDZP basis set provides an optimal balance, offering near-triple-ζ accuracy at double-ζ cost with minimal BSSE [6]. When highest accuracy for noncovalent interactions is required, aug-cc-pVTZ represents the practical sweet spot, sufficiently reducing BSIE/BSSE without prohibitive cost [43]. For exploratory studies on large systems, def2-SVP offers reasonable performance, though researchers should acknowledge its limitations for quantitative predictions [6].
Error Correction Protocol Recommendations
For geometry optimizations of molecular complexes, CP corrections should be applied throughout the optimization process, as BSSE significantly affects equilibrium structures, particularly with smaller basis sets [14]. For single-point energy calculations with triple-ζ basis sets or larger, CP corrections become less critical, as natural BSSE reduction occurs through basis set completeness [43]. When working with wavefunction-based methods like CCSD(T), employing explicitly correlated F12 corrections with appropriate basis sets (e.g., cc-pVXZ-F12) dramatically reduces BSIE, potentially obviating the need for separate CP corrections [36].
Basis set selection remains a critical consideration in quantum chemical applications to drug discovery, directly impacting both computational cost and result accuracy. The evolving landscape of basis set development, exemplified by specialized sets like vDZP and advanced strategies like FNOs, continues to push the Pareto frontier of efficiency. By strategically matching basis sets and error correction protocols to specific research objectives—prioritizing different approaches for geometry optimizations versus single-point energies, or for covalent versus noncovalent interactions—researchers can significantly enhance productivity without compromising scientific rigor. As quantum computational methods advance, intelligent basis set selection will grow increasingly important for harnessing these technologies in practical drug discovery applications.
Mitigating the Central Atom Bias in CP Corrections for Large, Asymmetric Systems
The accurate calculation of interaction energies in large, asymmetric molecular systems represents a significant challenge in computational chemistry, particularly for applications in drug development and materials science. The Basis Set Superposition Error (BSSE) arises from the use of incomplete basis sets in quantum chemical calculations, leading to exaggerated binding energies for molecular complexes [7]. This error is especially problematic in large systems with significant size asymmetry between components, where the assignment of atoms to fragments becomes ambiguous. The standard approach for mitigating this error, the Counterpoise (CP) correction method developed by Boys and Bernardi, calculates interaction energies by recomputing monomer energies using the entire basis set of the complex [7]. However, this method faces fundamental conceptual challenges when applied to large, asymmetric systems, particularly those involving chemical reactions or significant charge transfer. This guide provides a comprehensive comparison between the traditional CP correction and the alternative Chemical Hamiltonian Approach (CHA), evaluating their performance, limitations, and applicability for contemporary research challenges in drug development and molecular sciences.
Theoretical Frameworks: CP Correction vs. Chemical Hamiltonian Approach
Counterpoise Correction: Methodology and Limitations
The Counterpoise method operates on a simple principle: for every geometrical arrangement of a molecular complex, the energy of each free monomer is recalculated using the entire basis set of the supermolecule. This theoretically eliminates the artificial stabilization caused by monomers "borrowing" basis functions from their partners [7]. The standard CP correction protocol involves:
- Geometry Optimization: Optimize the geometry of the complex at the desired level of theory.
- Single-Point Energy Calculation: Compute the total energy of the complex, E_AB.
- Fragment Calculations in Supermolecule Basis: Calculate the energies of individual fragments (EA and EB) using the full basis set of the complex, with ghost atoms occupying the positions of the partner fragment.
- Interaction Energy Calculation: Determine the CP-corrected interaction energy as ΔECP = EAB - (EA + EB).
Despite its widespread use, CP correction faces significant challenges in large, asymmetric systems:
- Central Atom Bias: In asymmetric systems, the CP correction becomes direction-dependent. For a reaction A + B → C, calculating the correction with respect to reactants versus products yields different barrier heights [7]. This creates an inherent ambiguity in reaction barrier calculations.
- Fragment Assignment Ambiguity: Near transition states or in systems with significant delocalization, atoms cannot be clearly assigned to specific fragments. The original CP method, designed for two fragments, cannot adequately handle systems that naturally decompose into three or more components [7].
- Additivity Assumption: The CP method assumes BSSE effects and true intermolecular interactions are additive, which may not hold in systems with strong electronic coupling [7].
Chemical Hamiltonian Approach: Theoretical Foundation
The Chemical Hamiltonian Approach offers an alternative theoretical framework that addresses BSSE at a more fundamental level. Instead of applying posteriori corrections to calculated energies, CHA identifies and excludes the terms in the Hamiltonian responsible for BSSE [7]. The methodology involves:
- Hamiltonian Modification: The electronic Hamiltonian is modified to eliminate terms that lead to non-physical charge delocalization between fragments.
- Direct Calculation: Interaction energies are calculated directly from the modified Hamiltonian without need for fragment recalculations.
- Size Consistency: The approach naturally maintains size consistency across different system sizes and configurations.
Experience shows that CHA and CP methods typically yield similar results for standard intermolecular interaction energies when significant BSSE is present in uncorrected calculations [7]. However, the CHA formalism has not yet been fully developed for chemical reactions, limiting its application in studying reaction mechanisms [7].
Comparative Performance Analysis
Quantitative Comparison in Model Systems
Table 1: Performance Comparison of BSSE Correction Methods for Ion-Water Clusters
| Method | System Type | Error Trend with System Size | Fragment Assignment | Reaction Application |
|---|---|---|---|---|
| CP Correction | F⁻(H₂O)₁₅ clusters | Runaway error accumulation for semilocal DFT [46] | Ambiguous for ≥3 fragments [7] | Problematic for transition states [7] |
| CHA | Weakly bonded complexes | Comparable to CP for intermolecular interactions [7] | Well-defined | Not yet developed [7] |
| Uncorrected DFT | Small water clusters (H₂O)₆ | Minor effects [46] | Not applicable | Applicable but inaccurate |
Table 2: Performance of CP Correction with Different Density Functionals
| Functional Type | Example Functionals | MBE(n) Convergence | Mitigation Strategies |
|---|---|---|---|
| Semilocal GGA | PBE | Wild oscillations, divergent [46] | Energy-based screening [46] |
| Hybrid (<50% exact exchange) | B3LYP, PBE0 | Significant oscillations [46] | ≥50% exact exchange [46] |
| Meta-GGA | ωB97X-V, SCAN, SCAN0 | Insufficient elimination of divergent behavior [46] | Not effective with continuum models [46] |
| Hartree-Fock | - | Convergent behavior [46] | - |
Case Study: The PH₃ + H → PH₂ + H₂ Reaction
Research on the model reaction PH₃ + H → PH₂ + H₂ highlights the fundamental challenges of CP corrections in chemical reactions. When applying CP correction to the transition state, the treatment becomes direction-dependent [7]. Calculating the correction with respect to reactants (PH₃ + H) versus products (PH₂ + H₂) yields different barrier heights, making the reaction barrier ill-defined [7]. This ambiguity is particularly severe for asymmetric reactions, while symmetric reactions (e.g., HO + HOH → HOH + OH) show less directional dependence [7].
The study concluded that there is "no perfect and practicable solution" to the problem of BSSE correction for bimolecular reactions, requiring researchers to select an appropriate compromise based on their specific system [7]. This fundamental limitation persists in current computational chemistry approaches for reaction modeling.
Delocalization Error in Many-Body Expansion
Recent research reveals that delocalization error interacts with the many-body expansion to create a feedback loop leading to runaway error accumulation, particularly in ion-water clusters [46]. For F⁻(H₂O)₁₅ clusters, PBE-based MBE(n) calculations show wild oscillations that worsen as n increases, making the expansion effectively divergent [46]. This problematic behavior is most pronounced for semilocal functionals but remains serious for hybrid functionals and meta-GGAs.
The combinatorial increase in the number of n-body terms drives this divergence despite individual ΔE_IJ⋯ corrections decreasing order-by-order [46]. This effect is dramatically exacerbated by the presence of anions, with fluoride-containing subsystems contributing disproportionately to the error accumulation [46].
Advanced Methodologies for Specific Scenarios
Generalized CP Schemes for Multi-Component Systems
For systems that naturally decompose into three or more fragments, such as transition states where transferred atoms cannot be assigned to either reactant, researchers have proposed generalized CP schemes:
- Turi-Dannenberg Scheme: The energy of each monomer is computed using the entire supermolecule basis [7].
- Hierarchical Scheme: The total energy is decomposed into 1-body, 2-body, and higher contributions with separate CP corrections calculated for each component [7].
These approaches aim to address the fragment assignment ambiguity but increase computational complexity and may not be universally applicable.
Modern Computational Strategies for Large Systems
Table 3: Advanced Computational Methods for Large System Calculations
| Method | Principle | Application Scope | BSSE Treatment |
|---|---|---|---|
| QM/MM | Quantum region embedded in molecular mechanics field | Biological systems (1000s of atoms) [47] | CP for QM region only |
| Method of Increments | Many-body expansion with localized orbitals | Solids and surfaces [47] | Component-specific corrections |
| SCC-DFTB | Self-consistent charge density-functional tight-binding | Biological molecules [47] | Parametric, no explicit BSSE |
| Plane-Wave DFT | Periodic boundary conditions with plane-wave basis | Molecules on surfaces, nanoclusters [47] | No BSSE but other limitations |
Mitigation Strategies for Delocalization Error
Research on F⁻(H₂O)N clusters demonstrates that several strategies can help mitigate but not eliminate the divergent behavior in DFT-based MBE calculations:
- High Exact Exchange: Hybrid functionals with ≥50% exact exchange can counteract but not fully eliminate problematic oscillations [46].
- Energy-Based Screening: Culling unimportant subsystems based on energy thresholds can successfully forestall divergent behavior [46].
- Hartree-Fock Reference: HF calculations show convergent MBE(n) behavior but lack electron correlation effects [46].
- Alternative Boundary Conditions: Dielectric continuum boundary conditions and density correction approaches show limited effectiveness [46].
The Scientist's Toolkit: Essential Computational Reagents
Table 4: Research Reagent Solutions for BSSE-Corrected Calculations
| Tool Category | Specific Tools | Function | Applicability |
|---|---|---|---|
| Electronic Structure Codes | GAUSSIAN, Q-CHEM, FRAGME∩T | Perform CP-corrected calculations [7] [46] | Standard quantum chemistry applications |
| Correction Protocols | Standard CP, Generalized CP (Turi-Dannenberg) | Implement BSSE correction schemes [7] | Direction-dependent and multi-fragment systems |
| Fragment-Based Approaches | Many-Body Expansion, Method of Increments | Enable large-system calculations [47] [46] | Solids, surfaces, and extended systems |
| Wavefunction Analysis | Localized Orbital Analysis, Charge Decomposition | Identify fragment assignment ambiguity [7] | Transition states and delocalized systems |
| Error Mitigation | Energy-Based Screening, High-Exact-Exchange Functionals | Control error accumulation in fragment methods [46] | Large, asymmetric ion-containing systems |
The mitigation of central atom bias in CP corrections remains an unresolved challenge in computational chemistry, particularly for large, asymmetric systems relevant to drug development. The traditional CP correction method, while robust for standard intermolecular complexes, shows fundamental limitations for reaction modeling and large systems with delocalized electronic structures. The Chemical Hamiltonian Approach offers theoretical advantages but has not been developed for chemical reactions, limiting its practical utility [7].
For researchers tackling these challenges, we recommend:
- For Reaction Modeling: Acknowledge the inherent ambiguity in CP-corrected barrier heights and report both reactant- and product-corrected values for asymmetric reactions [7].
- For Large Systems with Ions: Implement energy-based screening to control error accumulation in fragment-based methods and consider hybrid functionals with high exact exchange [46].
- For Method Development: Prioritize approaches that explicitly address the fragment assignment problem in transition states and highly delocalized systems.
The field requires continued development of fundamentally new approaches that move beyond the limitations of current BSSE correction schemes, particularly for the complex, asymmetric systems that dominate modern chemical and pharmaceutical research.
Optimizing Workflows for Large Biomolecules and High-Throughput Virtual Screening
High-throughput virtual screening (HTVS) has become an indispensable tool in modern computational chemistry and drug discovery, enabling researchers to rapidly evaluate millions or even billions of molecular candidates for desired properties. However, the enormous search space containing the candidates and the substantial computational cost of high-fidelity property prediction models make screening practically challenging [48]. The central challenge lies in optimizing these workflows to maximize both accuracy and efficiency—a balance that requires sophisticated computational strategies and careful attention to methodological details. This challenge is particularly acute when dealing with large biomolecules, where accurate energy calculations are essential for reliable results.
Within this context, the accurate calculation of interaction energies faces a fundamental theoretical hurdle: the basis set superposition error (BSSE). This error arises from the use of finite basis sets in quantum chemical calculations, where the description of intramolecular energy improves within molecular complexes because basis orbitals of partner molecules become available [7]. The most commonly used cure for BSSE is the counterpoise (CP) correction according to the Boys and Bernardi scheme, which recalculates the energy of free molecules using the whole basis of the complex [7]. However, this approach faces significant conceptual and practical difficulties when applied to reacting systems and transition states, where the assignment of atoms to specific molecular fragments becomes ambiguous [7].
This review examines current methodologies for optimizing HTVS workflows, with particular emphasis on the interplay between accuracy in energy calculations and efficiency in screening large compound libraries. We compare the predominant counterpoise correction method with alternative approaches and provide experimental data on their performance in real-world screening scenarios.
Theoretical Foundations: BSSE Correction Methods
Counterpoise Correction Approach
The counterpoise (CP) correction method, initially proposed by Boys and Bernardi, remains the most widely used approach for addressing BSSE in computational chemistry [7]. In this approach, interaction energy is determined by recomputing the energy of the free molecules for every geometrical arrangement of the system using the whole basis of the complex. This method operates on the assumption of additivity between BSSE effects and true intermolecular interactions, an approximation that generally performs well for simple intermolecular complexes [7].
However, the application of CP correction becomes problematic when studying chemical reactions, particularly in the transition state region. As noted in foundational research, "the atom (or group of atoms) which is transferred from one molecule to another cannot legitimately be assigned to either of them" in the neighborhood of the transition state [7]. This ambiguity leads to mathematical inconsistencies where the barrier height becomes ill-defined, as the corrections calculated in forward and backward directions generally differ [7].
Chemical Hamiltonian Approach
The Chemical Hamiltonian Approach (CHA) presents an alternative theoretical framework for addressing BSSE. Instead of applying corrections to monomer energies, CHA modifies the calculation of the overall system by identifying and excluding those terms of the Hamiltonian which cause BSSE [7]. Experience shows that CHA and CP methods typically yield similar results for intermolecular interaction energies, even when uncorrected calculations show substantial BSSE [7].
A significant limitation, however, is that "the CHA formalism has not yet been developed for reactions" [7], restricting its application primarily to non-covalent interactions. This represents a substantial gap in methodology for studying chemical reactions involving large biomolecules.
Extension to Multi-Fragment Systems
For complex molecular systems involving more than two fragments, both CP and CHA face theoretical challenges. Two generalized CP schemes have been proposed for such systems: the Turi and Dannenberg approach, where each monomer's energy is computed in the whole supermolecule basis, and the hierarchical scheme of White and Davidson and Valiron and Mayer for N-component systems, where total energy is decomposed into 1-body, 2-body, etc., energy contributions with separate CP corrections for each component [7].
Table 1: Comparison of BSSE Correction Methods
| Method | Theoretical Basis | Applicability | Limitations |
|---|---|---|---|
| Counterpoise Correction | Recomputes monomer energies in supermolecule basis | Intermolecular complexes, some reaction systems | Ambiguous fragment assignment in transition states; direction-dependent results |
| Chemical Hamiltonian Approach | Excludes BSSE-causing terms from Hamiltonian | Intermolecular complexes | Not developed for chemical reactions |
| Generalized CP Schemes | Extends CP to multiple fragments with hierarchical decomposition | Multi-fragment systems | Increased computational complexity; implementation challenges |
Modern High-Throughput Virtual Screening Workflows
The Efficiency Challenge in Large-Scale Screening
The exponential growth of virtual chemical libraries has created unprecedented computational challenges for HTVS. Where earlier libraries contained hundreds of thousands of compounds, modern libraries like ZINC have grown from 700k to 120 million structures between 2005 and 2015, and now contain roughly 1 billion molecules [49]. Some non-enumerated libraries implicitly define up to 10¹⁰ to 10²⁰ possible compounds [49]. Exhaustively screening libraries of this magnitude requires computational resources inaccessible to most researchers—one cited study required 475 CPU-years for a single screening campaign [49].
Bayesian Optimization for HTVS
Bayesian optimization has emerged as a powerful framework for addressing the computational cost of large-scale virtual screening. This approach uses a surrogate model trained on previously acquired data to guide the selection of subsequent experiments, dramatically reducing the number of calculations required [49]. Instead of screening every molecule in a library indiscriminately, Bayesian optimization iteratively selects the most promising candidates for evaluation based on predictions from the surrogate model.
In practice, this method can identify 94.8% or 89.3% of the top-50,000 ligands in a 100-million member library after testing only 2.4% of candidate ligands using specific acquisition strategies [49]. The enrichment factor (EF)—defined as the ratio of the percentage of top-k scores found by model-guided search to percentage found by random search—can reach 11.9 for some model configurations, representing nearly an order-of-magnitude improvement in efficiency [49].
Multi-Fidelity Screening Pipelines
An alternative optimization approach involves constructing HTVS pipelines that consist of multi-fidelity models, optimally allocating computational resources to models with varying costs and accuracy to maximize return on computational investment [50] [48]. This framework enables an adaptive operational strategy where researchers can trade accuracy for efficiency depending on their specific needs [48].
In such pipelines, inexpensive but approximate filters rapidly eliminate unpromising candidates, while progressively more accurate—and computationally expensive—methods evaluate the remaining compounds. This tiered approach ensures that computational resources are concentrated on the most promising regions of chemical space.
Experimental Protocols and Performance Comparison
Implementation of Active Learning for Docking
Schrödinger's Therapeutics Group has developed a sophisticated implementation of active learning for docking screens through their Active Learning Glide (AL-Glide) technology. This approach combines machine learning with docking to enable enrichment with docking for libraries of billions of compounds [51]. The protocol follows these specific steps:
- Initial Batch Selection: A manageable batch of compounds is selected from the complete dataset and docked using traditional methods.
- Model Training: These docked compounds are added to the training set for a machine learning model.
- Iterative Refinement: The process iterates as the ML model becomes an increasingly accurate proxy for the docking method.
- Full Library Evaluation: The trained ML model rapidly evaluates the entire library, identifying the most promising candidates.
- Final Verification: Traditional docking calculations are performed on the best-scoring compounds identified by the model [51].
This implementation "dramatically narrows down the number of compounds made or purchased and assayed in the lab—frequently achieving double-digit hit rates" compared to traditional virtual screening approaches that typically yield 1-2% hit rates [51].
Performance Comparison of Optimization Methods
Experimental evaluations of different Bayesian optimization parameters provide quantitative data on their relative performance:
Table 2: Performance of Bayesian Optimization Configurations for Virtual Screening
| Surrogate Model | Acquisition Function | Top-100 Hits Found | Enrichment Factor | Computational Cost |
|---|---|---|---|---|
| Random Forest (RF) | Greedy | 51.6% (±5.9) | 9.2 | 6% of library screened |
| Random Forest (RF) | Upper Confidence Bound | Similar to Greedy (EF=7.7) | 7.7 | 6% of library screened |
| Neural Network (NN) | Greedy | 66.8% | 11.9 | 6% of library screened |
| Message Passing Neural Network | Upper Confidence Bound | 94.8% of top-50k in 100M library | N/A | 2.4% of library screened |
The performance comparison reveals that "using a directed-message passing neural network we can identify 94.8% or 89.3% of the top-50,000 ligands in a 100M member library after testing only 2.4% of candidate ligands" [49], demonstrating the remarkable efficiency gains possible with optimized workflows.
Integrated Workflows for Hit Identification
Modern virtual screening workflows often integrate multiple techniques in a sequential fashion. Schrödinger's approach exemplifies this integrated methodology:
- Ultra-large scale screening with AL-Glide to efficiently explore billions of compounds.
- Rescoring of top candidates using more sophisticated docking programs like Glide WS that leverage explicit water information.
- Absolute Binding Free Energy calculations (ABFEP+) for accurate scoring of diverse chemotypes.
- Active learning for ABFEP+ to further optimize the use of this computationally expensive method [51].
This integrated approach has been successfully applied to a broad range of targets across multiple screening campaigns, achieving "double-digit hit rates" consistently [51].
The Scientist's Toolkit: Essential Research Reagents and Software
Table 3: Essential Software Tools for Biomolecular Simulation and Virtual Screening
| Tool Name | Type | Primary Applications | Key Features |
|---|---|---|---|
| GROMACS | Open-source MD software | Biomolecular simulations, especially proteins and nucleic acids | Highly efficient parallel computing and GPU acceleration [52] |
| LAMMPS | Open-source MD software | Materials science, nanotechnology, complex physical systems | High customizability; supports millions of particles [52] |
| AMBER | Commercial/Free | Biomolecular simulations, drug design | Optimized for protein folding and small molecule binding [52] |
| CHARMM | Commercial | Biomolecular simulations | Specializes in long-range interactions [52] |
| NAMD | Commercial | Large biomolecular complexes | Highly parallelized for large-scale systems [52] |
| AutoDock Vina | Open-source | Molecular docking | Fast, accurate docking scoring functions [49] |
| AlphaFold2 | Free | Protein structure prediction | Neural network-based structure prediction [53] |
| MolPAL | Open-source | Bayesian optimization for screening | Implements multiple acquisition functions and surrogate models [49] |
Optimizing workflows for large biomolecules and high-throughput virtual screening requires careful attention to both theoretical accuracy and computational efficiency. The persistent challenge of BSSE correction, particularly the limitations of both counterpoise correction and Chemical Hamiltonian approaches for reaction systems, highlights the need for continued methodological development [7]. Meanwhile, modern screening workflows have dramatically improved efficiency through Bayesian optimization and active learning approaches, reducing computational requirements by over an order of magnitude while maintaining high hit identification rates [49] [51].
The most successful implementations combine multiple strategies: multi-fidelity modeling to allocate computational resources efficiently [50] [48], machine learning-guided docking to prioritize promising compounds [51], and sophisticated free energy calculations for final verification [51]. These approaches enable researchers to navigate the vast chemical space of modern billion-compound libraries while managing computational costs. As virtual libraries continue to grow, further development of these optimized workflows will be essential for maintaining the pace of discovery in computational chemistry and drug development.
Benchmarking Accuracy: A Rigorous Comparison of CP and CHA Performance
In computational chemistry, the Basis Set Superposition Error (BSSE) is a fundamental challenge that arises when calculating interaction energies between molecular fragments using the supermolecular approach. This error originates from the use of finite basis sets, where fragments artificially lower their energy by partially using the basis functions of neighboring fragments, leading to an overestimation of binding energies. Two primary methodologies have been developed to correct for this error: the Counterpoise (CP) correction proposed by Boys and Bernardi, and the Chemical Hamiltonian Approach (CHA) developed by Mayer [14]. While both aim to provide accurate interaction energies, their theoretical foundations and practical implementations differ significantly. The CP approach is a posteriori correction, whereas CHA is an a priori method that modifies the Hamiltonian itself. Understanding their relative performance across different system types is crucial for researchers in computational chemistry, drug design, and materials science who rely on accurate molecular interaction data.
Theoretical Foundations of CP and CHA
The Counterpoise (CP) Correction Method
The CP correction method, introduced by Boys and Bernardi, employs a simple but effective strategy to estimate BSSE. It calculates the interaction energy as the difference between the energy of the supermolecule and the energies of individual fragments, all computed with the full basis set of the complex. The standard CP-corrected interaction energy, ΔEc(fCP), is given by:
ΔEc(fCP) = EABAB(AB) - EABAB(A) - EABAB(B) [14]
Here, EABAB(A) represents the energy of fragment A computed with its own basis set plus the basis functions of fragment B at the geometry of the complex (often referred to as "ghost orbitals"). This approach effectively eliminates the artificial stabilization caused by the borrowing of basis functions between fragments. The CP method has gained widespread adoption due to its conceptual simplicity and ease of implementation in quantum chemistry software packages. However, it does involve additional computational cost, as it requires separate calculations for each fragment with the complete basis set of the complex.
The Chemical Hamiltonian Approach (CHA)
In contrast to the a posteriori nature of CP correction, the Chemical Hamiltonian Approach (CHA) developed by Mayer provides an a priori solution to BSSE. CHA modifies the fundamental Hamiltonian of the system to explicitly exclude the effects of basis set superposition from the outset. This method projects out the overcompleteness introduced by the overlapping basis sets of different fragments, resulting in a modified Hamiltonian that is theoretically free from BSSE. Despite their conceptual differences, studies have indicated that both CP and CHA methodologies tend to yield similar numerical results for many molecular systems [14]. However, CHA has been applied less frequently than CP in practical computational studies, potentially due to its more complex theoretical foundation and implementation requirements in standard quantum chemistry packages.
Experimental Protocols for Method Comparison
Computational Setup and System Selection
A systematic investigation into the accuracy of CP versus CHA across different system types was conducted at the MP2 (Møller-Plesset second-order perturbation) level of theory using Gaussian 03 program [14]. The study employed three Pople-style basis sets: 6-311+G(d), 6-311+G(d,p), and 6-311++G(d,p), along with Dunning's correlation-consistent basis sets (aug-cc-pVnZ, where n = D, T, Q) to examine basis set convergence. The research focused on six hydrogen-bonded trimer complexes: (NH₃)₃, (H₂O)₃, (HF)₃, (PH₃)₃, (H₂S)₃, and (HCl)₃ [14]. This selection enabled a comprehensive analysis across different periodic elements and bonding characteristics.
The experimental workflow followed a rigorous multi-step process to ensure comparable and meaningful results between the two correction methods, as illustrated below:
Types of CP Corrections Evaluated
The study implemented two distinct types of CP corrections to thoroughly evaluate structural effects:
Type 1 Correction: This approach considered the trimer as a combination of three interacting monomers (A, B, and C). The CP correction was applied between the (A---B) dimer and the C monomer, as well as between the (B---C) dimer and the A monomer [14].
Type 2 Correction: This method treated the system as an interaction between two fragments, where one fragment was a dimer (A---B) and the other was a monomer (C) [14].
For each correction type, researchers analyzed changes in intermolecular distances, interaction energies, and the relative contribution of BSSE to the total interaction energy. This multifaceted approach provided insights into how different fragmentation schemes affect the predicted molecular properties.
Comparative Performance Analysis Across Systems
Quantitative Comparison of Structural Parameters
The systematic investigation revealed how CP corrections affect optimized geometries across different trimer complexes. The table below summarizes the key findings for period 2 element trimers:
Table 1: CP Correction Effects on Trimers of Period 2 Elements [14]
| Trimer System | Basis Set | Distance Changes with CP Correction | %δE(fCP) | Remarks |
|---|---|---|---|---|
| (NH₃)₃ | 6-311+G(d) | R₁, R₂, R₃ all lengthened | 19.3% | Consistent lengthening observed |
| (NH₃)₃ | aug-cc-pVQZ | Minimal lengthening | <5% | Basis set convergence reduces CP effect |
| (H₂O)₃ | 6-311+G(d) | All distances lengthened | 22.3% | Similar pattern to ammonia trimer |
| (HF)₃ | 6-311+G(d) | Bond lengths increased | 27.3% | Highest %δE among period 2 trimers |
For trimers composed of period 3 elements ((PH₃)₃, (H₂S)₃, (HCl)₃), the CP corrections showed similar trends but with varying magnitudes. The (HCl)₃ system exhibited the most pronounced effects, with %δE(fCP) values reaching up to 48.6% with the 6-311+G(d) basis set [14]. This indicates that weaker binding complexes experience relatively larger BSSE contributions compared to their stronger-binding counterparts.
Accuracy Comparison Between CP and CHA
While both methods generally provide similar results, the comparative analysis revealed nuanced differences in their performance:
Table 2: Direct Comparison of CP vs. CHA Performance Characteristics [14]
| Performance Metric | CP Correction | CHA Approach | Practical Implications |
|---|---|---|---|
| Theoretical Foundation | A posteriori correction | A priori Hamiltonian modification | CHA more theoretically rigorous |
| Implementation Complexity | Low | Moderate | CP more widely adopted |
| Computational Cost | Moderate (extra fragment calculations) | Low to moderate | CHA potentially more efficient |
| Structural Effects | Lengthened intermolecular distances | Similar structural changes | Comparable predictions |
| Basis Set Dependence | High for small basis sets | Similar dependence | Both methods converge with large basis sets |
| Reported Accuracy | High agreement with CHA | High agreement with CP | Essentially equivalent results |
The study concluded that despite their conceptual differences, both methodologies tend to yield similar numerical results [14]. The relative contribution of BSSE correction, %δE(fCP), remained significant even with large basis sets like aug-cc-pVQZ, emphasizing the importance of these corrections for chemical accuracy regardless of the chosen method.
Essential Research Reagent Solutions
To implement the methodologies described in this comparison, researchers require specific computational tools and approaches. The table below outlines the essential "research reagents" for conducting such studies:
Table 3: Essential Research Reagents for BSSE Correction Studies
| Research Reagent | Function/Description | Example Applications |
|---|---|---|
| Quantum Chemistry Software | Platforms implementing CP and CHA methods | Gaussian 03, other packages with BSSE correction options [14] |
| Electron Correlation Methods | Account for electron-electron interactions | MP2, CCSD(T), QCISD(T) for accurate interaction energies [14] |
| Basis Sets | Mathematical functions for electron orbitals | Pople-style (6-311G), Dunning cc-pVnZ series [14] |
| Molecular Model Systems | Well-characterized complexes for benchmarking | Hydrogen-bonded trimers, van der Waals complexes [14] |
| Geometry Optimization Algorithms | Locate minimum energy structures | Gradient-based methods for efficient convergence [14] |
| Frequency Analysis Tools | Verify stationary points and compute ZPVE | Harmonic approximation for vibrational analysis [14] |
Implications for Research and Development
The comparative analysis between CP and CHA corrections has significant implications for computational drug development and materials design. In molecular docking studies, accurate prediction of protein-ligand binding energies is paramount for virtual screening campaigns. The findings demonstrate that BSSE corrections are particularly crucial for weakly interacting systems, which are common in drug-receptor interactions. The research shows that with medium-sized basis sets, BSSE can account for 20-50% of the calculated interaction energy in hydrogen-bonded systems [14], highlighting the potential for significant errors in uncorrected calculations.
For researchers working with non-covalent interactions in drug design, such as π-π stacking, hydrogen bonding, and van der Waals forces, these results underscore the importance of consistently applying BSSE corrections. The essential equivalence of CP and CHA results provides flexibility in method selection, with CP often being the practical choice due to its wider implementation in computational chemistry software. As the field moves toward high-throughput virtual screening and machine learning-based drug discovery, understanding these fundamental corrections ensures that training data and predictive models are based on physically meaningful interaction energies rather than computational artifacts.
This comprehensive comparison between CP and CHA corrections across different system types reveals that both methods provide essentially equivalent results for molecular interaction energies and optimized geometries [14]. The key finding is that despite their different theoretical foundations, both approaches successfully correct for BSSE and yield comparable predictions when applied consistently. The research demonstrates that CP corrections systematically lengthen intermolecular distances across all studied trimer complexes, with the magnitude of correction being highly dependent on the basis set quality.
Future research directions should explore the performance of these corrections in more complex chemical systems, including transition metal complexes, supramolecular assemblies, and biological macromolecules. Additionally, with the increasing importance of quantum chemical calculations in machine learning potentials and materials informatics, understanding the transferability of these corrections to emerging methodologies will be crucial. As computational chemistry continues to play a pivotal role in drug development and materials design, the rigorous treatment of BSSE remains an essential consideration for achieving chemical accuracy in molecular simulations.
In quantum chemistry, the pursuit of accurate molecular simulations is perpetually balanced against computational feasibility. Finite basis sets, used to approximate molecular orbitals, introduce two key errors: Basis Set Superposition Error (BSSE), which artificially lowers interaction energies as fragments borrow each other's basis functions, and Basis Set Incompleteness Error (BSIE), which arises from an insufficient description of the electron correlation cusp and short-range interactions [1] [54]. To mitigate BSSE, two primary correction schemes are employed: the a posteriori Counterpoise (CP) correction and the a priori Chemical Hamiltonian Approach (CHA).
This guide objectively compares the performance of these corrections, detailing how their efficacy and accuracy evolve as basis sets increase in size and quality. Understanding this convergence is critical for researchers making informed decisions in fields like drug development, where accurate prediction of non-covalent interactions is paramount [55] [56].
Theoretical Foundations of BSSE Corrections
The Counterpoise (CP) Correction
The Counterpoise method is a corrective procedure applied after an initial calculation. It estimates the BSSE by performing additional "ghost" calculations where each monomer's energy is recalculated using the full, supermolecular basis set, but with the other fragment's atoms represented by only their basis functions (without atomic nuclei or electrons) [1]. The BSSE is then subtracted from the uncorrected interaction energy. A noted limitation is its potential for inconsistent effects across different areas of a potential energy surface [1].
The Chemical Hamiltonian Approach (CHA)
In contrast, the CHA seeks to prevent BSSE from the outset. It modifies the Hamiltonian operator itself by removing terms that would allow for basis set mixing between monomers [1]. This creates a computational framework that is inherently free from BSSE, without the need for a separate correction step. It has been argued that this method treats all fragments more equally compared to the CP method [1].
Comparative Analysis: Correction Efficacy Across Basis Sets
The performance of BSSE corrections is intrinsically linked to the size and diffuseness of the atomic orbital basis set. The table below summarizes the general evolution of correction efficacy.
Table 1: Evolution of BSSE Correction Efficacy with Increasing Basis Set Size
| Basis Set Tier | Typical Examples | BSSE Severity (Uncorrected) | CP Correction Efficacy | CHA Correction Efficacy | Typical Application Context |
|---|---|---|---|---|---|
| Double-ζ (DZ) | def2-SVP, cc-pVDZ | Large, significant | Moderate; residual error can be substantial [6] | Moderate; prevents mixing but BSIE is large | Preliminary screening; cost-effective studies [6] |
| Triple-ζ (TZ) | def2-TZVP, cc-pVTZ | Moderate | Good; often sufficient for qualitative accuracy | Good; balanced performance | Standard for reliable single-point energies [6] |
| Augmented Triple-ζ | def2-TZVPPD, aug-cc-pVTZ | Reduced but non-negligible | High; crucial for accurate non-covalent interactions [55] | High; effective for accurate interaction energies [1] | Benchmarking non-covalent interactions [55] |
| Quadruple-ζ (QZ) & Larger | aug-cc-pVQZ, aV5Z | Small | Very High; errors disappear rapidly [1] | Very High; errors disappear rapidly [1] | High-accuracy thermochemistry; approaching CBS limit [57] |
Key Trends and Research Insights
- Rapid Error Disappearance: The inherent BSSE and the residual errors from both CP and CHA corrections disappear more rapidly than the total BSSE itself as the basis set is enlarged [1]. In very large basis sets, the corrected and uncorrected energies converge.
- The Diffuse Function Conundrum: Diffuse functions (e.g., in
aug-basis sets) are essential for accurately modeling non-covalent interactions, dispersion forces, and anions (the "blessing of accuracy") [55]. However, they drastically reduce the sparsity of the one-particle density matrix, increasing computational cost and worsening the "curse of sparsity" [55]. This makes BSSE corrections not just beneficial, but often necessary when using these basis sets. - Comparative Performance: While the CP and CHA are conceptually different, they often yield similar numerical results [1]. Some studies suggest the CP correction can be less consistent because central atoms have greater freedom to mix with all available ghost functions, whereas CHA treats all fragments equally [1].
Experimental Protocols for Benchmarking
To objectively compare the performance of BSSE corrections, researchers typically employ a standardized workflow. The following diagram visualizes this process.
Diagram 1: Workflow for benchmarking BSSE correction efficacy across basis sets.
Detailed Methodology
- System Selection: Choose a model system with well-characterized non-covalent interactions, such as a DNA base pair or a water dimer [55].
- Geometry Optimization: Optimize the geometry of the complex and its monomers using a high-level method (e.g., CCSD(T)) with a large, diffuse basis set to minimize uncertainty in the structure [57].
- Single-Point Energy Calculations:
- Perform energy calculations for the complex and monomers across a series of basis sets (e.g., cc-pVDZ → aug-cc-pV5Z).
- For each basis set, calculate the uncorrected interaction energy.
- Apply the CP correction using the standard ghost-atom protocol [1].
- Perform calculations using the CHA to obtain inherently corrected energies [1].
- Reference Data: Use experimental binding energies or highly accurate theoretical values (e.g., from CCSD(T) at the Complete Basis Set (CBS) limit or explicitly correlated F12 calculations) as a benchmark [57].
- Error Analysis: Compute the deviation of the CP-corrected, CHA-corrected, and uncorrected interaction energies from the reference value for each basis set. Plotting these deviations reveals the convergence profile.
Table 2: Essential Computational Reagents and Resources
| Tool/Resource | Type | Primary Function | Relevance to BSSE Studies |
|---|---|---|---|
| Basis Set Exchange [55] | Software Database | Provides access to hundreds of published Gaussian basis sets. | Crucial for obtaining consistent, well-documented basis sets across the DZ-to-QZ spectrum for systematic studies. |
| Correlation-Consistent (cc-pVXZ) [58] | Basis Set Family | Systematically convergent basis sets designed for correlation energy recovery. | The gold standard for performing basis set convergence studies and extrapolating to the CBS limit. |
| Karlsruhe (def2-XVP) [55] | Basis Set Family | Efficient, segmented contracted basis sets for elements across the periodic table. | Widely used in production and benchmarking calculations due to their good cost-to-accuracy ratio. |
| vDZP [6] | Specialized Basis Set | A compact double-ζ basis set designed to minimize BSSE and BSIE. | Useful as a low-cost option for screening; demonstrates that targeted basis set design can reduce reliance on corrections. |
| GMTKN55 [6] | Benchmark Database | A comprehensive collection of 55 benchmark sets for main-group thermochemistry and non-covalent interactions. | Provides a standardized test suite for rigorously evaluating the performance of methods and corrections. |
| Explicitly Correlated (F12) Methods [57] | Computational Method | Accelerates basis set convergence by explicitly including the interelectronic distance r12. | Not a BSSE correction, but a powerful alternative for reaching CBS-quality energies with smaller basis sets, thereby side-stepping the BSSE problem. |
The efficacy of BSSE correction methods is inextricably linked to basis set size. For small double-ζ basis sets, BSSE is large and corrections, while beneficial, still leave significant residual error. The use of diffuse functions in triple-ζ and larger basis sets is often mandatory for chemical accuracy in non-covalent interactions, making rigorous BSSE correction equally mandatory. Both the Counterpoise and Chemical Hamiltonian Approach show rapidly improving efficacy with larger basis sets, with their results typically converging. For the highest accuracy, as in drug development studies targeting protein-ligand binding, employing large, augmented quadruple-ζ basis sets with robust BSSE correction remains the most reliable path to definitive results [56].
The accurate prediction of molecular interactions is a cornerstone of modern computational drug discovery. This guide focuses on the performance of advanced computational methods, evaluated within the critical context of two major therapeutic target classes: kinase inhibitors and metalloenzyme complexes. The accurate modeling of these systems presents significant challenges for computational chemistry, particularly concerning the treatment of non-covalent interactions, transition metal centers, and basis set superposition error (BSSE). Within this framework, research into BSSE correction methods, specifically the counterpoise (CP) correction and the chemical Hamiltonian approach (CHA), is highly relevant. These methods aim to correct the artificial stabilization introduced by the use of incomplete basis sets in quantum mechanical calculations, a factor that can significantly impact the accuracy of computed binding energies in drug-relevant systems [59] [60]. This guide objectively compares the application and outcomes of various computational protocols by presenting experimental data and case studies from the scientific literature, providing a practical resource for researchers.
Computational Methods and Challenges in Drug Discovery
The computational modeling of drug-relevant systems requires a multi-faceted approach, leveraging different methods based on the specific question being investigated. The following table summarizes the key quantum mechanical methods, their optimal applications, and their inherent limitations, particularly for complex biological systems.
Table 1: Key Computational Methods in Drug Discovery
| Method | Strengths | Limitations | Best Applications in Drug Discovery | Computational Scaling |
|---|---|---|---|---|
| Density Functional Theory (DFT) | High accuracy for ground states; handles electron correlation; wide applicability [59] [60] | Accuracy depends on exchange-correlation functional; struggles with large biomolecules [59] [60] | Binding energies, electronic properties, reaction mechanisms, metalloenzyme inhibitor design [59] [60] | O(N³) |
| Hartree-Fock (HF) | Fast convergence; reliable baseline; well-established theory [59] [60] | Neglects electron correlation; poor for weak interactions (e.g., van der Waals) [59] [60] | Initial geometry optimization, charge distribution calculation [59] [60] | O(N⁴) |
| Quantum Mechanics/Molecular Mechanics (QM/MM) | Combines QM accuracy with MM efficiency; handles large biomolecules [59] [60] | Complex setup at the QM/MM boundary; method-dependent accuracy [59] [60] | Enzyme catalysis, protein-ligand interaction studies [59] [60] | O(N³) for QM region |
A primary challenge in using these methods, especially for calculating interaction energies in protein-ligand complexes, is Basis Set Superposition Error (BSSE). BSSE is an artificial lowering of energy that occurs when using finite basis sets, making interactions appear stronger than they are. The two main approaches to correct for BSSE are:
- Counterpoise (CP) Correction: A post-hoc correction that calculates the interaction energy using a "ghost" basis set to account for the partner molecule's orbitals.
- Chemical Hamiltonian Approach (CHA): An a priori method that projects out the overlapping basis functions during the calculation itself.
The performance and accuracy of these approaches are a subject of ongoing research, particularly in systems like metalloenzymes where metal-ligand coordination is critical.
Case Study 1: Metalloenzyme Inhibitors
Metalloenzymes, which constitute nearly half of all enzymes, are central to many diseases, making them attractive therapeutic targets [61] [62]. A key strategy in inhibiting these enzymes involves designing small molecules that coordinate the active site metal ion via a metal-binding pharmacophore (MBP) [61] [63].
Performance Data and Selectivity Profiling
The following table summarizes experimental inhibition data for a selection of clinically approved and well-characterized metalloenzyme inhibitors, showcasing their potency and the diversity of MBPs used.
Table 2: Experimental Inhibition Data for Selected Metalloenzyme Inhibitors
| Inhibitor | Target Protein | IC₅₀ (nM) | Metal-Binding Group (MBP) | Therapeutic Indication | FDA Approved |
|---|---|---|---|---|---|
| Acetazolamide [63] | Human Carbonic Anhydrase II (hCAII) | 25 | Sulfonamide | Glaucoma | Yes |
| Captopril [63] | Angiotensin Converting Enzyme (ACE) | 21 | Thiol | Hypertension | Yes |
| SAHA (Vorinostat) [63] | Histone Deacetylase (HDAC) | 10 | Hydroxamic Acid | Cutaneous T-cell lymphoma | Yes |
| CGS 27023A [63] | MMP-1, MMP-3, MMP-9 | 8 - 43 | Hydroxamic Acid | Investigational (Cancer) | No |
| 1,2-HOPO-2 [63] | MMP-12 | 18 | Hydroxypyridinone | Investigational (Cancer) | No |
Despite concerns about promiscuity, rigorous selectivity profiling has demonstrated that metalloenzyme inhibitors can be highly selective for their intended targets. A systematic study evaluating inhibitors with diverse MBGs (hydroxamic acid, carboxylate, thiol, etc.) against a panel of metalloenzymes (CA, MMPs, ACE, HDAC, tyrosinase) found minimal cross-inhibition, indicating that the drug-like "backbone" of the molecule confers significant specificity [63]. Furthermore, these inhibitors were unable to remove Fe³⁺ from the transport protein transferrin, even at high concentrations, alleviating concerns about systemic metal ion depletion [63].
Detailed Experimental Protocol: Selectivity Profiling
Objective: To evaluate the selectivity of metalloenzyme inhibitors and their potential to chelate metal ions from biological proteins [63]. Materials:
- Inhibitors: A panel of compounds representing diverse MBGs (e.g., hydroxamic acid, carboxylate, thiol, sulfonamide).
- Enzymes: A panel of purified metalloenzymes (e.g., hCAII, MMP-2, MMP-9, MMP-12, ACE, HDAC-2, Tyrosinase).
- Assay Reagents: Specific fluorogenic or colorimetric substrates for each enzyme, appropriate reaction buffers.
- Metal Chelation Assay: Holo-transferrin (iron-bound), buffer. Methodology:
- Enzyme Inhibition Assay:
- For each enzyme, prepare a series of inhibitor concentrations in a suitable buffer.
- Incubate the enzyme with the inhibitor for a defined period (e.g., 15-30 minutes).
- Initiate the reaction by adding the enzyme-specific substrate.
- Monitor the reaction progress (e.g., fluorescence or absorbance) in real-time.
- Calculate the % inhibition for each concentration and determine IC₅₀ values.
- Metal Ion Removal Assay:
- Incubate holo-transferrin with a high concentration of the inhibitor (e.g., 1 mM) for a defined period.
- Use spectroscopic methods or competition assays with chelating agents to detect the release of free Fe³⁺.
- Compare against a positive control (e.g., deferoxamine) and a negative control (buffer alone).
Research Reagent Solutions
Table 3: Essential Research Reagents for Metalloenzyme Studies
| Reagent / Resource | Function and Application | Example / Specification |
|---|---|---|
| Metal-Binding Fragment (MBF) Libraries [62] | Provides a diverse set of chelating chemotypes for screening against metalloenzyme targets to identify novel hits. | Custom libraries based on picolinic acid isosteres or generalized property characterization [62]. |
| Computational Docking Software | Predicts the binding pose and affinity of inhibitors within the enzyme's active site. | Requires customization for metal ions (e.g., parameterization, specific scoring functions) [62]. |
| Quantum Chemical Descriptors [62] | Electronic and topological descriptors calculated by DFT improve QSAR model performance for predicting inhibitor activity. | Ionization potential, electron affinity, atomic charges, multipole moments [62]. |
Case Study 2: Kinase Inhibitors
Kinases represent a major drug target class, with small-molecule kinase inhibitors (SMKIs) constituting a significant portion of FDA-approved drugs [61]. While most approved kinase inhibitors do not directly engage active site Mg²⁺ ions [61], the accurate computational description of their binding, which often involves π-π stacking and hydrogen bonding in the hinge region, remains a rigorous test for QM methods.
Performance Data and Binding Mode Analysis
c-Met kinase, a high-value target in oncology, illustrates the evolution of inhibitor design from Type II to more selective Type I inhibitors. A recent study combined scaffold hopping and structure-guided synthesis to develop novel 1,3,4-thiadiazolo[2,3-c]-1,2,4-triazin-4-one derivatives as Type I c-Met inhibitors [64].
Table 4: Experimental c-Met Kinase Inhibition Data for Novel Thiadiazolo-Triazinone Derivatives
| Compound | c-Met Inhibition Rate at 10 µM | IC₅₀ (nM) | Binding Mode |
|---|---|---|---|
| 3d [64] | > 80% | Not fully reported | Type I |
| 5d [64] | > 80% | Promising (specific value not listed) | Type I |
| 5f [64] | > 80% | Promising (specific value not listed) | Type I |
| Lead Compound (5d) [64] | N/A | Exhibited the best anti-proliferative activity | Type I (U-shaped conformation) |
Molecular docking simulations revealed that these inhibitors adopt a characteristic U-shaped conformation within the ATP-binding pocket, a hallmark of Type I inhibitors. Key interactions include π-π stacking with Tyr 1230 and hydrogen bonds with Met 1160 in the hinge region, which are critical for their potency and selectivity [64].
Detailed Experimental Protocol: Structure-Guided Kinase Inhibitor Design
Objective: To design and synthesize novel, selective Type I c-Met kinase inhibitors using a scaffold-hopping and structure-guided approach [64]. Materials:
- Software: Molecular Operating Environment (MOE) for docking simulations.
- Protein Structure: PDB code 3CCN (c-Met kinase).
- Chemistry: Reagents and solvents for organic synthesis (e.g., from Merck, Sigma-Aldrich); thin-layer chromatography (TLC) for reaction monitoring; silica gel for column chromatography.
- Assay: c-Met kinase activity assay kit. Methodology:
- Molecular Docking:
- Prepare the protein structure (3CCN) by removing water molecules and adding hydrogen atoms.
- Dock proposed candidate molecules into the c-Met kinase binding site.
- Analyze the binding poses to confirm adoption of the U-shaped Type I binding mode and key interactions (π-π stacking with Tyr 1230, H-bonds with Met 1160).
- Chemical Synthesis:
- Based on docking results, select candidate structures for synthesis.
- Perform multi-step synthesis as described, using TLC to monitor reaction progress.
- Purify final compounds using techniques like column chromatography and recrystallization.
- Confirm structure and purity using melting point determination, NMR, and mass spectrometry.
- Biological Evaluation:
- Test synthesized compounds for c-Met kinase inhibition at a single concentration (e.g., 10 µM) to determine inhibition rate.
- For hits showing >80% inhibition, determine full IC₅₀ values using a dose-response curve.
- Evaluate the anti-proliferative activity of the most potent inhibitors against c-Met over-expressed cancer cell lines.
Research Reagent Solutions
Table 5: Essential Research Reagents for Kinase Inhibitor Studies
| Reagent / Resource | Function and Application | Example / Specification |
|---|---|---|
| Kinase Assay Kits | Measure the inhibitory activity of novel compounds against purified kinase targets. | c-Met kinase activity assay. |
| Molecular Docking Software | Utilized for structure-based design and predicting binding modes of inhibitors. | MOE (Molecular Operating Environment) [64]. |
| Chemical Synthesis Reagents | High-purity starting materials and solvents for the synthesis of candidate inhibitor molecules. | Commercial suppliers (e.g., Merck, Sigma-Aldrich) [64]. |
The case studies presented here demonstrate the critical role of rigorous experimental validation in computational drug discovery. For metalloenzyme inhibitors, selectivity profiling against diverse enzyme panels is essential to counter concerns about promiscuity, revealing that well-designed inhibitors can achieve high specificity [63]. In kinase inhibitor development, combining computational docking with synthetic chemistry and biological testing enables the rational design of compounds with desired binding modes and improved selectivity profiles [64]. The performance of any computational method, including CP and CHA for BSSE correction, must ultimately be validated against such experimental data. The continued integration of accurate quantum mechanical methods, systematic experimental protocols, and structural insights is key to advancing the discovery of effective therapeutics for complex targets.
The accurate computation of energy and molecular properties is foundational to advancements in fields ranging from drug development to materials science. A significant source of error in quantum chemical calculations, particularly for weakly interacting systems, is the Basis Set Superposition Error (BSSE). This error arises from the use of finite basis sets, where the description of a monomer in a complex is artificially improved by "borrowing" basis functions from its interacting partner, leading to overestimated binding energies [7]. To mitigate BSSE, two principal theoretical approaches have been developed: the Counterpoise (CP) correction method and the Chemical Hamiltonian Approach (CHA). The CP method, introduced by Boys and Bernardi, corrects for BSSE by recalculating the energy of each isolated monomer using the entire basis set of the complex, thereby estimating the magnitude of the error [7] [65]. In contrast, the CHA is an a priori method that identifies and eliminates the terms in the Hamiltonian responsible for BSSE, preventing the error from occurring in the first place [66]. This guide provides a quantitative comparison of these two methodologies, evaluating their accuracy, computational cost, and applicability in modern research, with a special focus on their performance within Density Functional Theory (DFT) and for modeling chemical reactions.
Theoretical Frameworks and Computational Protocols
The Counterpoise (CP) Correction Method
The standard Counterpoise (CP) correction procedure for a dimer system (AB) is a widely used a posteriori method to correct for BSSE. The protocol involves several key steps. First, a geometry optimization of the dimer (AB) is performed at a chosen level of theory. The single-point energy of the dimer, E(AB), is then computed in its own basis set. Finally, the single-point energies of the isolated monomers A and B are calculated, but each is computed using the full, entire basis set of the dimer (AB). The CP-corrected interaction energy is subsequently calculated using the formula: ΔE_CP = E(AB) - [E(A in AB basis) + E(B in AB basis)] [7] [65]. This method can be generalized beyond two-body systems. For more complex systems, such as trimers, a hierarchical scheme can be applied. This involves recursively calculating BSSE-corrected energy terms (one-body, two-body, three-body, etc.) by recomputing each energy contribution in the basis sets of increasingly larger subclusters [66].
The Chemical Hamiltonian Approach (CHA)
The Chemical Hamiltonian Approach (CHA) offers a fundamentally different, a priori strategy. Instead of correcting energies after the calculation, CHA modifies the Hamiltonian operator itself to remove the terms that are the physical origin of BSSE [66]. The core of the CHA methodology involves a systematic identification and elimination of the BSSE-inducing terms within the Hamiltonian, which correspond to the unphysical "borrowing" of basis functions. The electronic structure calculation, such as a Hartree-Fock or DFT computation, is then performed using this corrected, BSSE-free Hamiltonian. Experience shows that the CHA and CP methods typically yield very close results for standard intermolecular interaction energies, even when the uncorrected results show significant BSSE [7] [66].
Comparative Accuracy Analysis: Performance Across Systems
The performance of CP and CHA methods varies significantly depending on the chemical system and property being investigated. The following table summarizes their comparative performance based on key studies.
Table 1: Comparative Accuracy of CP and CHA Methods
| System / Property | Counterpoise (CP) Performance | Chemical Hamiltonian Approach (CHA) Performance | Key Comparative Findings |
|---|---|---|---|
| Intermolecular Complexes (e.g., Water Dimer) | Good performance; reliable interaction energies when applied correctly [7]. | Excellent performance; results are closely aligned with CP-corrected values [66]. | Both methods effectively reduce BSSE, yielding comparable results for non-covalent interactions [7] [66]. |
| Reaction Pathways & TS | Problematic; suffers from direction-dependence in asymmetric reactions, leading to ill-defined barrier heights [7]. | Not yet developed for chemical reactions, representing a significant limitation [7]. | CP has conceptual flaws for TS; CHA is currently inapplicable, leaving a critical methodological gap. |
| Many-Body Systems | Can be generalized via hierarchical schemes for N-body interactions [66]. | Theoretical framework is general, but practical implementations are less common. | Both can be extended in theory, but CP's hierarchical scheme has been more directly demonstrated [66]. |
| Integration with DFT | Widely implemented and standard in most quantum chemistry codes [6]. | Has been successfully implemented within DFT using Gaussian basis sets [66]. | CHA performs as well within DFT as it does for Hartree-Fock methods [66]. |
The Critical Challenge of Transition States and Reaction Paths
A particularly difficult area for BSSE correction is the modeling of chemical reaction pathways, especially the transition state (TS). The CP method faces two major problems in this context. First, for an asymmetric bimolecular reaction (A + B → C + D), the CP correction at the TS depends on the choice of fragments. Correcting with respect to the reactant molecules (A and B) yields a different barrier height than correcting with respect to the product molecules (C and D). This direction-dependence makes the barrier height ill-defined [7]. Second, at the TS, the transferred atom(s) cannot be legitimately assigned to one fragment or the other. This necessitates treating the system as consisting of three or more fragments, for which the standard two-fragment CP method is inadequate [7]. While generalized CP schemes for N-component systems have been proposed, they are more complex and less commonly used [7] [66]. Notably, the CHA formalism has not yet been extended to handle chemical reactions, which currently limits its utility in this critical area of research [7].
Workflow and Decision Pathways for Researchers
The following diagram illustrates the logical decision process a researcher should follow when selecting a BSSE correction method for a given study, based on the system type and the research objective.
Essential Research Reagent Solutions
The experimental and computational research in this field relies on a suite of software, basis sets, and model systems. The following table details these key "research reagents" and their functions.
Table 2: Key Research Reagents and Computational Tools
| Reagent / Tool | Type | Primary Function in BSSE Research |
|---|---|---|
| libint | Software Library | Computes one- and two-electron integrals, a foundational component for building new quantum chemistry codes [32]. |
| libxc | Software Library | Provides a standardized, comprehensive collection of exchange-correlation functionals for DFT calculations [32]. |
| vDZP Basis Set | Basis Set | A purpose-made double-zeta basis set designed to minimize BSSE, enabling efficient and accurate calculations with various density functionals [6]. |
| GMTKN55 Database | Benchmark Database | A comprehensive collection of 55 benchmark sets used to quantify the accuracy of quantum chemical methods, including main-group thermochemistry, non-covalent interactions, and barrier heights [6]. |
| ωB97X-3c | Composite DFT Method | A composite method combining a functional, the vDZP basis set, and empirical corrections, exemplifying the modern approach to achieving high accuracy with low computational cost [6]. |
| Model Reaction: PH₃ + H → PH₂ + H₂ | Chemical System | A well-studied model reaction used to illustrate the conceptual and numerical difficulties of applying BSSE corrections to reaction barriers [7]. |
The statistical evaluation of errors in energy and molecular properties reveals that both the Counterpoise correction and the Chemical Hamiltonian Approach are highly effective for mitigating BSSE in standard intermolecular complexes, with both methods yielding quantitatively similar results. However, for the critical task of modeling chemical reaction pathways and transition states, significant challenges remain. The CP method becomes conceptually ill-defined for asymmetric reactions, while the CHA is not yet formulated for this purpose. Future research should prioritize the extension of the CHA to reactive systems and the development of more robust, generalized CP schemes. Furthermore, the integration of these methods with modern, efficient computational frameworks—such as the use of BSSE-minimizing basis sets like vDZP in high-throughput workflows—will be essential for improving the accuracy and reliability of quantum chemical calculations in drug development and materials science.
Accurately simulating molecular systems is a cornerstone of computational chemistry, with direct implications for drug discovery and materials science. Achieving high fidelity in these simulations often requires careful method selection to balance computational cost with physical accuracy. Two philosophical approaches emerge: one focuses on correcting specific errors in standard methods, such as the Counterpoise (CP) correction for the Basis Set Superposition Error (BSSF), while the other seeks to reformulate the underlying quantum mechanical model, as seen in the Chemical Hamiltonian Approach (CHA). This guide objectively compares these strategies, providing a framework for researchers to determine the optimal application for each method within the broader context of electronic structure calculation research.
Theoretical Foundations and Methodologies
The fundamental goal of many quantum chemistry simulations is to solve the electronic Schrödinger equation for molecules, a task that grows exponentially in complexity with system size [67]. The Hamiltonian operator is central to this endeavor, as it describes the system's energy and dynamics [67].
1. Counterpoise (CP) Correction: An A Posteriori Correction The CP method is not a standalone computational model but a corrective protocol applied to the results of standard quantum chemistry calculations like Hartree-Fock or Density Functional Theory (DFT). It specifically targets the Basis Set Superposition Error (BSSE), an artificial lowering of energy that occurs when fragmented parts of a system (e.g., two interacting molecules) use each other's basis functions to describe their own electrons. The CP correction involves calculating the energy of each fragment in the full, super-system basis set, thereby estimating and subtracting the BSSE from the total interaction energy.
2. Chemical Hamiltonian Approach (CHA): An A Priori Formulation In contrast, the CHA is a fundamental reformulation of the quantum chemical Hamiltonian. It is designed from the outset to be BSSE-free. The core idea is to rigorously separate the internal quantum mechanics of a fragment from the external potential provided by other fragments. This is achieved by redefining the Hamiltonian to exclude the possibility of a fragment's wavefunction being expanded on the basis functions of other fragments, thus preventing BSSE from arising in the first place.
The table below summarizes the core philosophical and practical differences between the two approaches.
Table 1: Fundamental Comparison of CP and CHA
| Feature | Counterpoise (CP) Correction | Chemical Hamiltonian Approach (CHA) |
|---|---|---|
| Core Philosophy | A posteriori error correction | A priori error-free formulation |
| Treatment of BSSE | Empirically estimates and subtracts the error | Formally prevents the error from occurring |
| Computational Overhead | Additional single-point energy calculations for fragments | Requires implementation of a non-standard Hamiltonian |
| Integration | Can be applied to most standard quantum chemistry methods | Requires specialized code and theoretical understanding |
| Interpretation | Yields BSSE-corrected interaction energies | Provides a BSSE-free potential energy surface from the start |
Comparative Analysis: Accuracy and Performance
Direct, side-by-side experimental comparisons of CP and CHA in recent literature are scarce, as the field has largely evolved toward benchmarking these methods against high-level reference data or testing them within novel computational frameworks. However, insights can be drawn from contemporary research on Hamiltonian-based simulations and error correction.
1. Application in Modern Hybrid and Machine Learning Models A significant trend in computational chemistry is integrating physical principles like Hamiltonians into machine learning (ML) models to improve their transferability and interpretability. For instance, one study developed a method that dynamically parameterizes a semiempirical Hamiltonian using a deep neural network (HIPNN+SEQM) [33]. This model, trained on small organic molecules, demonstrated remarkable accuracy when extended to much larger biochemical systems, successfully predicting energies, atomic forces, and other quantum chemical properties [33]. This highlights the value of a robust Hamiltonian-based foundation (conceptually aligned with CHA's philosophy) for achieving transferable accuracy across diverse chemical spaces.
Similarly, Hamiltonian simulation-based algorithms are being developed for quantum computers to tackle large-scale electronic structure problems. Methods like Hamiltonian simulation-based quantum-selected configuration interaction (HSB-QSCI) have shown promise for strongly correlated systems, capturing over 99% of correlation energies by efficiently sampling important electronic configurations [68].
2. Data Requirements and Computational Scalability The choice between a correction method and a reformed Hamiltonian can also be influenced by computational resources and system size.
Table 2: Practical Considerations for Method Selection
| Criterion | Counterpoise (CP) Correction | Chemical Hamiltonian Approach (CHA) |
|---|---|---|
| Best for System Size | Small to medium-sized non-covalent complexes | Systems where formal correctness of the potential energy surface is paramount |
| Data Requirements | Relies on the accuracy of the underlying base method (e.g., DFT) | Requires specialized implementation; modern ML hybrids need training data [33] |
| Handling of Strong Correlation | Limited by the base method's capabilities | Hamiltonian-based frameworks are often better suited for strong correlation [68] |
| Ease of Use | Widely available in standard software packages | Less common, requires specialized expertise or code |
The development of general neural network potentials (NNPs), such as EMFF-2025 for energetic materials, demonstrates a different path. These models achieve DFT-level accuracy for mechanical properties and decomposition behaviors by leveraging transfer learning, which reduces the need for massive new datasets [69]. This contrasts with the CP method, which does not learn from data but applies a fixed correction, and with the CHA, which is a first-principles approach.
Decision Framework: Selecting the Right Tool
The choice between CP and CHA is not a matter of which is universally better, but which is more appropriate for a specific research goal. The following workflow diagram outlines the key decision points for researchers.
Diagram 1: Decision workflow for CP vs. CHA
The Scientist's Toolkit: Essential Research Reagents
The following table details key computational "reagents" and tools essential for conducting research in this field.
Table 3: Key Research Reagent Solutions for Electronic Structure Studies
| Research Reagent | Function & Purpose |
|---|---|
| High-Performance Computing (HPC) Cluster | Provides the computational power for expensive quantum chemistry calculations (e.g., DFT, CCSD(T)) and neural network potential training [69]. |
| Quantum Chemistry Software | Platforms like Qiskit and PennyLane enable quantum algorithm simulation and ground state energy calculations, useful for benchmarking and developing new methods [70]. |
| Semiempirical Quantum Mechanics (SEQM) Packages | Software like PYSEQM provides fast, reduced-order Hamiltonian calculations that can serve as a base for more accurate ML-corrected or CHA-inspired models [33]. |
| Reference Datasets | High-quality data (e.g., molecular energies, forces from DFT) is crucial for training ML-potentials like HIPNN and for validating the accuracy of both CP and CHA methods [33] [69]. |
| Tensor Network Emulators | Tools like Fermioniq's Ava allow for the emulation of quantum algorithms on classical computers, facilitating the study of Hamiltonian simulation for larger systems than exact methods allow [67]. |
In the pursuit of chemical accuracy, both the Counterpoise correction and the Chemical Hamiltonian Approach offer distinct pathways. The CP correction remains a practical, widely accessible tool for refining interaction energy calculations in non-covalent complexes. In contrast, the CHA represents a more fundamental, formally rigorous solution for developing BSSE-free potential energy surfaces. The emerging paradigm, as seen in hybrid ML-Hamiltonian models, leans toward the CHA philosophy by building dynamically responsive and physically interpretable models from the ground up [33]. For researchers and drug development professionals, the optimal choice is guided by the problem at hand: prefer CP for efficient, post-hoc correction of standard method results, and lean toward CHA or its modern descendants when formal correctness, transferability, and a foundational understanding of the Hamiltonian are critical.
Conclusion
Both the Counterpoise correction and the Chemical Hamiltonian Approach provide essential pathways to mitigate BSSE, yet their performance is highly context-dependent. While CP is widely implemented, users must be cautious of its limitations, such as potential inconsistencies on energy surfaces and uneven error correction across atoms. CHA offers a more theoretically unified approach but may be less accessible in standard software. The choice between them should be guided by the specific molecular system, the property of interest, and the available computational resources. For the drug discovery community, the rigorous application of these corrections is not merely academic; it is a prerequisite for achieving chemical accuracy in predicting binding affinities, modeling reaction mechanisms, and ultimately accelerating the development of novel therapeutics. Future progress will likely involve the tighter integration of these corrections with emerging machine learning potentials and quantum computing methods to tackle ever-larger biological systems with confidence.
References
- 1. Basis set superposition error [https://en.wikipedia.org/wiki/Basis_set_superposition_error]
- 2. The Basis Set Superposition Error (BSSE) [https://zipse.cup.uni-muenchen.de/teaching/computational-chemistry-2/topics/the-basis-set-superposition-error-bsse/]
- 3. Optimizing the basis set extrapolation parameter for weak ... [https://www.sciencedirect.com/science/article/abs/pii/S2210271X25004311]
- 4. 7.8 Basis Set Superposition Error (BSSE) [https://manual.q-chem.com/5.0/sect0022.html]
- 5. Basis Set Superposition Error (BSSE) — ADF 2025.1 ... - SCM [https://www.scm.com/doc/ADF/Input/BSSE.html]
- 6. The vDZP Basis Set Is Effective For Many Density Functionals [https://www.rowansci.com/publications/vdzp]
- 7. Some difficulties in computing BSSE-corrected potential ... [https://www.sciencedirect.com/science/article/abs/pii/S0009261498011919]
- 8. Simulating the ghost: quantum dynamics of the solvated ... [https://www.nature.com/articles/s41467-021-20914-0]
- 9. At the Forefront of Medium-Sized Molecule Drug Research ... [https://www.chugai-pharm.co.jp/english/story/detail/20250817000001_48.html]
- 10. MCAT Intermolecular vs Intramolecular Forces Explained [https://www.inspiraadvantage.com/blog/mcat-intermolecular-vs-intramolecular-forces]
- 11. Visualizing chemical space networks with RDKit and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9798653/]
- 12. Drug Discovery Market Size Hit USD 158.74 Bn by 2034 [https://www.precedenceresearch.com/drug-discovery-market]
- 13. Impact of the Intramolecular Basis Set Superposition Error [https://pmc.ncbi.nlm.nih.gov/articles/PMC6832644/]
- 14. Effects of the basis set superposition error on optimized ... [https://www.sciencedirect.com/science/article/abs/pii/S0009261412008020]
- 15. Accurate proton affinity and gas-phase basicity values for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2970571/]
- 16. Computational investigation of the conformer selective ... [https://www.sciencedirect.com/science/article/abs/pii/S0022286022021202]
- 17. Conformational kinetics reveals affinities of protein ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4522757/]
- 18. Base-Pairing Energies of Proton-Bound Dimers and ... [https://link.springer.com/article/10.1007/s13361-015-1174-2]
- 19. BSSE-free SCF theories: a comment [https://www.sciencedirect.com/science/article/abs/pii/S0166128099004546]
- 20. Toward Real Chemical Accuracy on Current Quantum ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11137825/]
- 21. Estimating the Complete Basis Set Extrapolation Error ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12105030/]
- 22. 8.9 Ghost Atoms and Basis Set Superposition Error [https://manual.q-chem.com/5.3/chap_ecp.html]
- 23. 2.11. Counterpoise Corrections [https://orca-manual.mpi-muelheim.mpg.de/contents/essentialelements/counterpoise.html]
- 24. Extending quantum-mechanical benchmark accuracy to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12479982/]
- 25. Basis Set Superposition Corrections [https://psicode.org/psi4manual/master/nbody.html]
- 26. Behavior of counterpoise correction in many‐body ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9303541/]
- 27. Evaluating wavefunction methods, the counterpoise ... [https://pubmed.ncbi.nlm.nih.gov/40314268/]
- 28. A B3LYP study on counterpoise-corrected geometry ... [https://www.sciencedirect.com/science/article/abs/pii/S0166128009005685]
- 29. Computational insights into solvent encapsulation and host ... [https://www.nature.com/articles/s41598-025-20260-x]
- 30. Isotopic Consequences of Host–Guest Interactions [https://pmc.ncbi.nlm.nih.gov/articles/PMC8023698/]
- 31. A quantum chemical interaction energy dataset for ... [https://www.nature.com/articles/s41597-023-02443-1]
- 32. Open-Source Projects We Wish Existed - Rowan Scientific [https://rowansci.com/blog/open-source-projects-we-wish-existed]
- 33. Deep learning of dynamically responsive chemical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9271210/]
- 34. Better performance of Hartree–Fock over DFT: a quantum ... [https://link.springer.com/article/10.1007/s00894-023-05706-1]
- 35. The "Charlotte's Web" of Density-Functional Theory (DFT) [https://rowansci.com/publications/the-charlottes-web-of-dft]
- 36. Machine-learning interatomic potentials achieving CCSD(T ... [https://arxiv.org/html/2508.14306v1]
- 37. Benchmarking simulacra AI's Quantum Accurate Synthetic ... [https://arxiv.org/html/2511.07433v1]
- 38. (IUCr) Benchmarking quantum chemical methods with X-ray ... [https://journals.iucr.org/m/issues/2025/04/00/pl5046/]
- 39. Machine learning for accuracy in density functional ... [https://arxiv.org/html/2311.00196v2]
- 40. Can the Counterpoise Correction for Basis Set ... [https://pubmed.ncbi.nlm.nih.gov/26579908/]
- 41. Advancing Universal Deep Learning for Electronic ... [https://arxiv.org/html/2509.19877v2]
- 42. Improved sample-based quantum diagonalization via ... [https://research.ibm.com/publications/improved-sample-based-quantum-diagonalization-via-randomized-hamiltonian-simulation--1]
- 43. Basis Set Incompleteness Errors in Fixed-Node Diffusion ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12079793/]
- 44. Extending quantum-mechanical benchmark accuracy to ... [https://www.nature.com/articles/s41467-025-63587-9]
- 45. Improving the runtime of quantum phase estimation for ... [https://arxiv.org/html/2509.05733v1]
- 46. Delocalization error poisons the density-functional many-body ... [https://pubs.rsc.org/en/content/articlehtml/2024/sc/d4sc05955g]
- 47. Quantum Chemistry Methods for Large Molecular Systems [https://www.cecam.org/workshop-details/quantum-chemistry-methods-for-large-molecular-systems-554]
- 48. Optimizing high-throughput virtual screening campaigns [https://biomlsp.com/2023/11/05/optimizing-high-throughput-virtual-screening-campaigns/]
- 49. Accelerating high-throughput virtual screening through ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8188596/]
- 50. Optimal decision-making in high-throughput virtual ... [https://www.sciencedirect.com/science/article/pii/S2666389923002672]
- 51. Dramatically improving hit rates with a modern virtual ... [https://www.schrodinger.com/life-science/learn/white-papers/dramatically-improving-hit-rates-modern-virtual-screening-workflow/]
- 52. Top Molecular Dynamics Simulation Software Free, Open ... [https://www.iaanalysis.com/top-molecular-dynamics-simulation-software-open-source.html]
- 53. Comparison, Analysis, and Molecular Dynamics ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10525864/]
- 54. Correcting basis set incompleteness in wave function correlation ... [https://arxiv.org/html/2504.07192v1]
- 55. The Conundrum of Diffuse Basis Sets: A Blessing ... [https://arxiv.org/html/2502.04631v1]
- 56. Applications of density functional theory in COVID-19 drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8695517/]
- 57. The Impact of Larger Basis Sets and Explicitly Correlated ... [https://www.sciencedirect.com/science/article/abs/pii/S1574140016300019]
- 58. The behavior of density functionals with respect to basis set ... [https://pubmed.ncbi.nlm.nih.gov/15485223/]
- 59. Quantum Mechanics in Drug Discovery - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC12249871/]
- 60. Quantum Mechanics in Drug Discovery: A Comprehensive ... [https://www.mdpi.com/1422-0067/26/13/6325]
- 61. Targeting Metalloenzymes for Therapeutic Intervention - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6405328/]
- 62. Discovery of Novel Metalloenzyme Inhibitors Based on ... [https://www.mdpi.com/1420-3049/29/5/1096]
- 63. Investigating the Selectivity of Metalloenzyme Inhibitors [https://pmc.ncbi.nlm.nih.gov/articles/PMC3880651/]
- 64. Transforming Type II to Type I c-Met kinase inhibitors via ... [https://www.sciencedirect.com/science/article/abs/pii/S0045206821006817]
- 65. Energy-based measure of interaction strength [https://www.sciencedirect.com/science/article/abs/pii/B9780443334030000045]
- 66. The chemical Hamiltonian approach in density functional ... [https://www.sciencedirect.com/science/article/pii/0009261494012016]
- 67. Benefits and challenges of using tensor-network quantum ... [https://medium.com/@ellenmichael/benefits-and-challenges-of-using-tensor-network-quantum-emulators-for-hamiltonian-simulation-an-8452c09cb220]
- 68. Hamiltonian simulation-based quantum-selected ... [https://pubs.rsc.org/en/content/articlelanding/2025/cp/d5cp02202a]
- 69. EMFF-2025: a general neural network potential for ... [https://www.nature.com/articles/s41524-025-01809-w]
- 70. Advantages of two quantum programming platforms in ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01026-z]